Cells need to achieve very delicate balance between DNA synthesis and histone levels and ensure the synthesis of histone proteins just during the S phase where the DNA is being replicated. Restriction of histone biosynthesis to S phase is critical for maintenance of genomic stability and proper gene regulation. Many transcriptional and posttranscriptional factors contribute coordinately to regulate expression of histone proteins, including transcription of histone genes, efficiency of pre-mRNA processing (the percentage of mature mRNA that reach to cytoplasm), change in mRNA half- life and degradation of excess histone proteins. Stem-Loop Binding Protein (SLBP), which binds to 3' ends of histone mRNAs, is a key factor in histone biosynthesis. SLBP expression is cell cycle regulated without significant change in its mRNA level and this regulation is responsible for occurrence of bulk histone production during S phase. SLBP level is high during S phase and low in M and G1 phase until next S phase. It has been found that SLBP is degraded at the end of S phase due on double phosphorylation triggered by CyclinA/CDK2 and similarly in G1, the SLBP stability seems to be low. SLBP protein, which is not degraded at the end of S phase was found to be toxic for cells and effects the rate of DNA replication. These findings show that S phase limited expression of SLBP is very critical for histone mRNA biosynthesis.
Histone Biosynthesis and SLBP.
S phase is characterized with duplication of both DNA and histone proteins. Replication of eukaryotic chromosomes require synthesis of both DNA and sufficient histone proteins to package newly replicated DNA into chromatin structure. Histones are primary protein components of chromatins. They are involved in both DNA packaging and gene regulation. In metazoans majority of histones are canonical histone proteins and they are encoded by a family of replication dependent histone genes. These genes encode only four core histones: H2A, H2B, H3 and H4, all together make up nucleosome and H1 linker, which links nucleosomes (Ursprung, 1964). Synthesis of replication dependent histones is cell cycle regulated. In addition to canonical histones, there are several variant histones whose synthesis is not cell cycle regulated and they are different in mRNA structure from canonical ones. Replication-independent histone genes are constitutively expressed at low levels throughout the cell cycle.
Cells need to achieve very delicate balance between DNA synthesis and histone levels and ensure the synthesis of histone proteins just during the S phase where the DNA is being replicated. Restriction of histone biosynthesis to S phase is critical for maintenance of genomic stability and proper gene regulation. Many transcriptional and posttranscriptional factors contribute coordinately to regulate expression of histone proteins, including transcription of histone genes, efficiency of pre-mRNA processing (the percentage of mature mRNA that reach to cytoplasm), change in mRNA half- life and degradation of excess histone proteins (Marzluff et al., 2008)
One important feature of histone genes is that they are physically linked in a large cluster, enabling easier rapid transcription. Human replication-dependent histone genes are clustered on chromosomes 1 and 6. Six histone genes have been identified so far in the histone gene cluster at 1q21 (Braastad et al., 2004), while ∼50 histone genes have been found in the cluster at 6p21 (Albig and Doenecke, 1997; Marzluff et al., 2002). Genes encoding histones are constitutively transcribed by RNA polymerase II and their rate increases as cells approach S phase (DeLisle et al., 1983; R. Zhong et al., 1983). Transcription of histone genes are activated by Cyclin E-CDK2 dependent phosphorylation of NPAT in Cajal bodies. Depletion of NPAT results in substantial decrease in replication dependent histone RNA transcripts (X. Ye et al., 2003). Besides this, overexpression of NPAT promotes S phase entry, and coexpression of Cyclin E- Cdk2 enhances the effects of NPAT on cell cycle progression (A. Wang et al., 2004).
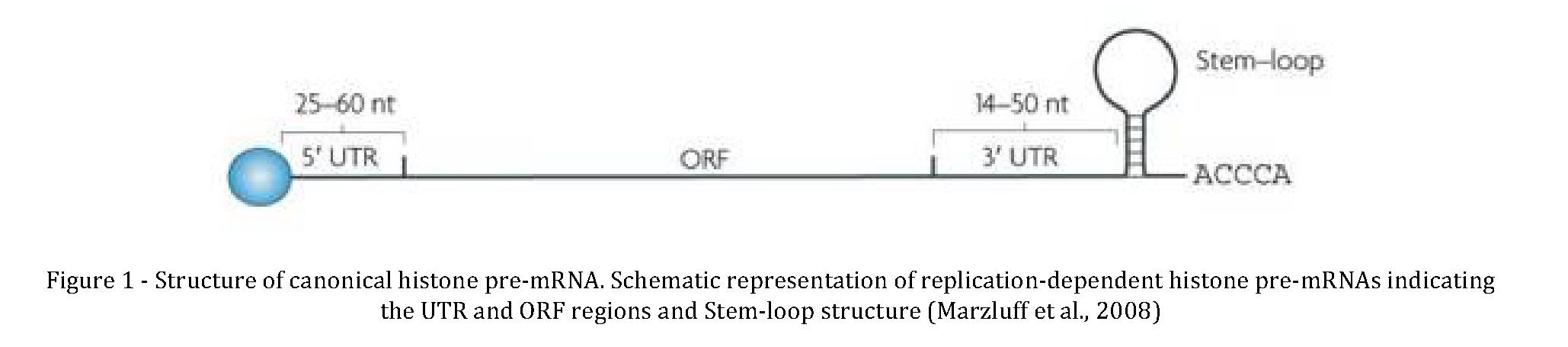
All five classes of histone proteins are encoded by so-called replication dependent histone mRNAs, which have unique and highly conserved stem loop at 3' end instead of poly-A tail (Figure 1.7). Like other mRNAs, replication dependent histone mRNAs have a 7-methyl-guanosine cap at 5' end. The 3' end of these mRNAs are required for histone stoichiometry in chromatin structure, as well as for regulation of the rate and timing of histone protein synthesis during cell cycle (Marzluff and Duronio, 2002; Marzluff et al., 2008). The expression histone mRNA is precisely controlled in a cell-cycle dependent manner with a precisely timed upregulation in S-phase when newly synthesized DNA hasto be packed into nucleosomes. At the beginning of S phase, the transcription of histone genes by RNA polymerase II increases from three to five fold whereas processing efficiency increases eight to ten fold, overall accounting for almost 35 fold increase in histone mRNA level (Marzluff, 2005). As cells exist S phase, the half-life and processing efficiency of histone mRNA is reduced, which together lead to instantaneous decrease in histone mRNA level (Harris et al., 1991).
Formation of canonical mature histone mRNA requires only endonucleatic cleavage at 3' end as they lack introns. For histone pre-mRNAs, cleavage occurs between the stem-loop and the histone downstream element (HDE), a purine rich sequence located about 15 nucleotides after the cleavage site (Dominski and Marzluff, 2007; Mowry and Steitz, 1987) and both these cisacting elements are required for proper processing reaction. A highly conserved 26 nucleotide stem-loop upstream of cleavage site sets up binding site for Stem-Loop Binding Protein or Hairpin Binding Protein (SLBP/HBP). SLBP is important for every aspects of histone mRNA metabolism and is the major player of processing that is cell cycle regulated protein parallel to histone mRNA (will be discussed in detail further) (Marzluff and Duronio, 2002).
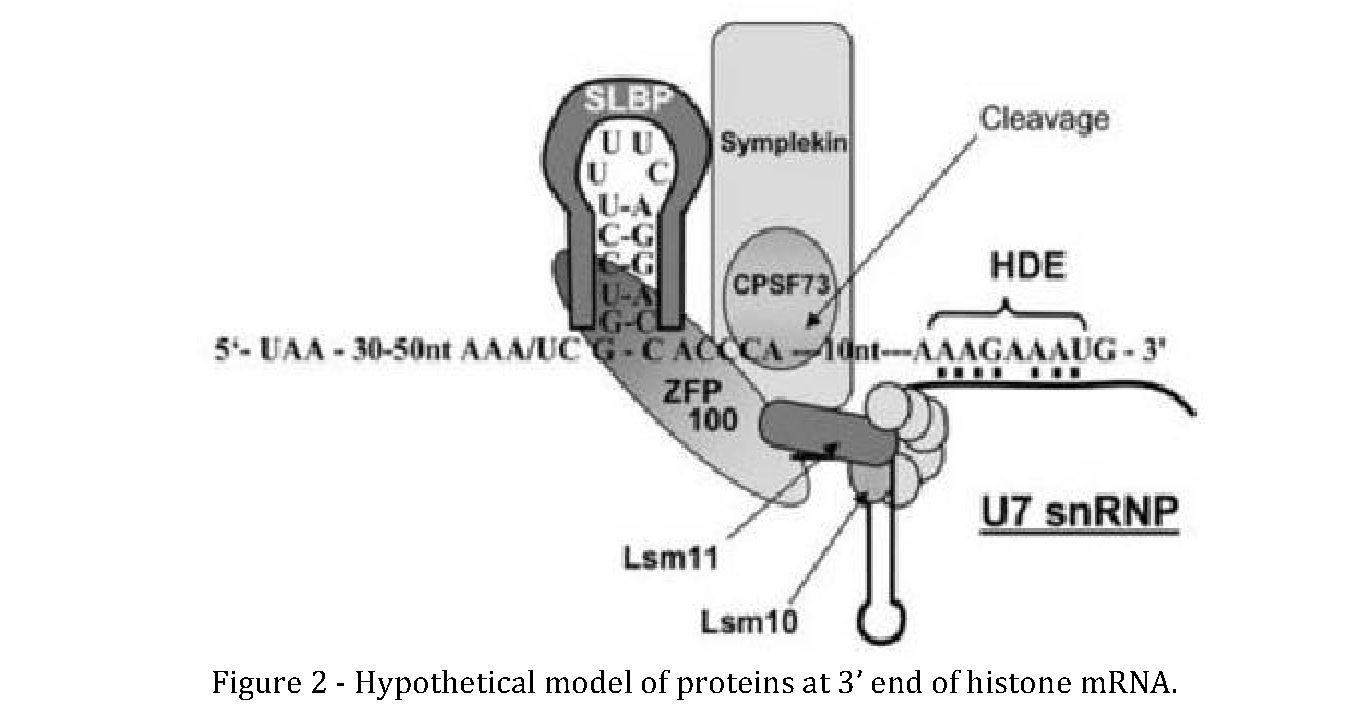
The second cis-acting element, so called HDE binds to RNA moiety of U7snRNP complex. U7snRNP is composed of 60 nucleotide U7snRNA and Sm ring. It is recruited to histone pre- mRNA primarily through formation of double stranded RNA between the 5' end of U7 snRNA and histone downstream element (HDE). Stem-Loop Binding Protein (SLBP) interacts with one of Sm ring proteins and stabilize the U7snRNP on the HDE, which is required for cleavage catalyzed by CPSF-73 (Dominski et al., 2005). Y** non-coding RNA is a medium sized ncRNA associates with CPSF complex and responsible for recruitment of other processing factor for enhanced processing efficiency of nascent histone mRNA (Kohn and Huttelmaier, 2016). Another recent study showed that, FUS, the RNA binding protein both in nucleus and cytoplasm, also associates with U7snRNP and nascent replication histone mRNAs predominantly during S phase. FUS is essential for proper processing of histone mRNA, as depletion of FUS leads to increased extended histone transcripts. FUS is also reported to bind to histone gene promoters and enhance binding of RNA polymerase II during S phase of cell cycle (Raczynska et al., 2015).
Unlike metazoan canonical histone mRNA, yeast histone mRNA lack Stem-Loop at 3' end and become polyadenylated like the rest of RNAs. It was reported that yeast histone mRNAs have shorter than average PolyA tails and the length of the PolyA tail varies during the cell cycle; S-phase histone mRNAs possess very short PolyA tails while in G1, the tail length is relatively longer. Polyadenylation cycle accounts for cell cycle regulation of histone production (Beggs et al., 2012). C.Elegans histone mRNA structure is similar to mammals, however no U7 homology was found. Instead, a small ncRNA at 3' UTR is responsible for efficient 3' end processing at S phase (Avgousti et al., 2012).
The illustration depicts possible interactions in the stable processing complex assembled on histone mRNA before cleavage reaction (Wagner and Marzluff, 2006). FLASH functions in histone gene transcription and has also been found to have a role in histone RNA processing. The FLASH-protein (FLICE-associated huge) associates with NPAT and is considered to be essential for connecting histone mRNA synthesis to 3'-end processing by recruitment of processing factors to nascent histone pre-mRNAs, thus reinforcing the formation of active HLBs (Histone Locus Bodies) (Yang et al., 2014; Yang et al., 2009). FLASH stimulates 3'-end processing in mammalian nuclear extracts, presumably via interaction with Lsm11 as observed in Drosophila (Dominski and Marzluff, 2007). FLASH also interacts with Ars2 (arsenite resistance protein 2) and knockdown of Ars2 inhibits 3'-end formation after the stem-loop and leads to end formation at downstream cryptic polyadenylation sites (Kiriyama et al., 2009). Lesion in this interaction triggers cell cycle arrest at S/G2.
After histone mature mRNA is produced, only SLBP remains bound to stem loop structure and escorts the mRNA from nucleus to cytoplasm. Histone mRNA is exported in TAP dependent manner in a very short time (Erkmann, Sanchez, et al., 2005; Schodhetman and Perry, 1972; K. D. Sullivan et al., 2009). Mammalian cells with knockdown SLBP fails to export processed histone mRNA to cytoplasm (Erkmann, Sanchez, et al., 2005; K. D. Sullivan et al., 2009) indicating that SLBP is required for this accompanying histone mRNA to cytoplasm.
3' end of histone mRNA is essential for translation in vivo (Gallie et al., 1996). SLBP is bound to histone mRNA throughout the transport and is required for its translation (Whitfield et al., 2004) and it is present in the cytoplasm only when histone mRNA is present (Erkmann, Sanchez, et al., 2005; Erkmann, Wagner, et al., 2005). SLBP in coordination with other proteins help to circularize histone mRNA to form ‘close-loop' conformation. This ensures mRNA protection and increase the efficiency of translation of histone mRNA by a mechanism similar to poly-A tailed mRNAs (Sanchez and Marzluff, 2002). Some of the mechanisms that mediate translation by SLBP have been revealed: SLBP interacts with SLIP1 that binds to E1F4G and plays role in translation of histone mRNA (Cakmakci et al., 2008). Inefficient translation triggers degradation of histone mRNA via association with Upf1 (Kaygun and Marzluff, 2005b).
The stem-loop at the 3' end of replication dependent histone mRNA is the cis element that mediates mRNA degradation. Rapid decay of histone mRNA requires SLBP, which is involved in recruiting the proteins necessary to add short oligo (U) tail to histone mRNA that is being translated (Mullen and Marzluff, 2008). Degradation of histone mRNAs requires Upf1 which binds to SLBP (Kaygun and Marzluff, 2005b), is a key regulator of the nonsense-mediated decay pathway, and ATR, a key regulator of the DNA damage checkpoint pathway activated during replication stress (Kaygun and Marzluff, 2005a). It is claimed that Pin1 mediated proline isomerization of SLBP triggers its removal from the stem-loop (Krishnan et al., 2012; M. Zhang et al., 2012)
by mediating dephosphorylation of SLBP on Thr 171 in RDB. It has also reported that phosphorylation of Thr 171 of SLBP is important for proper processing of histone mRNA (M. Zhang et al., 2012). Other than that phosphorylation of Ser 20 and Ser 23 by unknown kinases are proposed to be essential for SLBP degradation in nucleus (Krishnan et al., 2014). There is an additional evidence that points to importance of SLBP phosphorylation for processing via regulating its interaction with histone mRNA. SLBP in Drosophila (dSLBP) phosphorylation at C-terminal increases binding to mRNA and facilitate processing and as well as probably other aspects of histone mRNA metabolism (J. Zhang et al., 2014).
SLBP is required for all aspects of histone mRNA biosynthesis: proper processing of pre-mRNAs, nuclear export, translation and regulation of stability of mRNA as described above in detail. Mutation in stem loop that abolish the binding of SLBP results in no expression of processed histone mRNA in vivo (Pandey et al., 1994). To date, there is no evidence reported for functions of SLBP outside those directly involved in histone mRNA biosynthesis and this function is highly conserved among different species (Townley-Tilson et al., 2006). SLBP is required for cell cycle coupled histone mRNA production in many species (Arnold et al., 2008; E. Sullivan et al., 2001). Depletion of SLBP level in continuously cycling cells leads to accumulation of cells in S phase (Erkmann, Sanchez, et al., 2005; K. D. Sullivan et al., 2009; Zhao et al., 2004). One of the major defects caused in absence of SLBP is decrease in processing efficiency, which eventually leads to polyadenylated histone mRNAs formation, indicating that SLBP is required for proper pre-mRNA processing (Lanzotti et al., 2002; E. Sullivan et al., 2001). The loss of normal histone mRNA processing can result in the production of polyadenylated mRNAs from downstream polyA sites of the replication dependent histone genes (Narita et al., 2007; Wagner et al., 2007). The level of polyadenylated canonical histone mRNAs are very low in proliferating cells (Narita et al., 2007) but increase during tumorigenesis (Zhao et al., 2004). Expression of polyadenylated histonemRNAs have been revealed by microarray analyses in various sub-types of cancer (Zhao et al., 2004) . Polyadenylated histone mRNA are expressed in cell cycle independent manner and have distinct stability and translation regulations. Thus, they would be synthesized outside of S phase leading to genomic instability and alteration in gene regulations. All these evidences suggest that, SLBP has an essential role in restriction of histone biogenesis to S phase and contributes to maintenance of genomic stability.
SLBP is crucial for cell cycle regulation of histone mRNA.
Replication of eukaryotic chromosomes require synthesis of both DNA and sufficient histone proteins to package newly replicated DNA into chromatin structure. Cells adjust the level of histone mRNA to match the demand for histone protein which is dictated by the rate of DNA synthesis (reviewed in (Marzluff et al., 2008)). In mammalian cells there are four different regulatory mechanisms that contribute to proper histone protein level during the cell cycle: Transcription of histone genes, efficiency of pre-mRNA processing (the percentage of mature mRNA that reach to cytoplasm), change in mRNA half- life and degradation of excess histone proteins (Graves et al., 1987; Gunjan et al., 2006; Harris et al., 1991; Marzluff et al., 2008). Cells need to achieve very delicate balance between DNA synthesis and histone levels and ensure the synthesis of histone proteins occurs just during the S phase where the DNA is being replicated. Excess histones may alter the gene expression by nonspecifically binding to DNA (Au et al., 2008), cause mitotic chromosome loss (Meeks-Wagner and Hartwell, 1986) and mediate cytotoxicity by multiple mechanism (Singh et al., 2010). In contrary to this, scarcity of histone proteins results in cell death (Han et al., 1987). All five classes of histone proteins are encoded by so- called replication dependent histone mRNAs, which have unique and highly conserved stem loop at 3' end instead of poly-A tail (Busslinger et al., 1979; Connor et al., 1984). The 3'end of these mRNAs are required for histone stoichiometry in chromatin structure, regulation of the rate and timing of histone protein synthesis during cell cycle (Eckner et al., 1991; Marzluff and Duronio, 2002; Marzluff et al., 2008; Zanier et al., 2002). The level of histone mRNA increases dramatically as cells enter to S phase to cover the rapid need for new histone proteins.
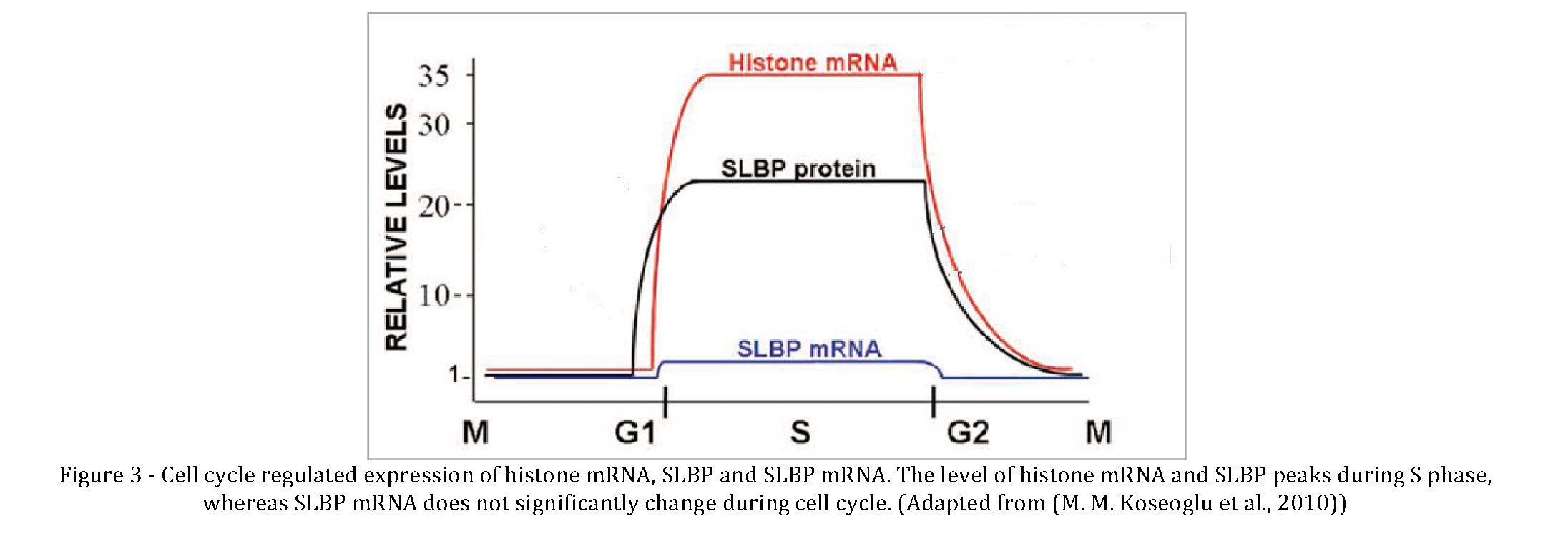
At the beginning of S phase, the transcription of histone genes by RNA polymerase II, increases from three to five fold whereas processing efficiency increases eight to ten fold, overall accounting for almost 35 fold increase in histone mRNA level (Marzluff, 2005). As cells exist S phase, the half-life and processing efficiency of histone mRNA is reduced, which together lead to instantaneous decrease in histone mRNA level (Marzluff, 2010). In mammals, the cell cycle regulation of SLBP (Stem Loop Binding Protein) that binds to stem loop at 3' end of histone mRNA accounts for the regulation of histone pre-mRNA processing (DeJong et al., 2002; Martin et al., 1997; Whitfield et al., 2000; Zheng et al., 2003). SLBP expression is cell cycle regulated in parallel to histone mRNA. The level of SLBP is very low in G1 and increases around 10 fold at the end of G1 phase where cells get ready for replication and it is degraded rapidly at the end of S phase. The amount of SLBP mRNA is almost stable during cell cycle, indicating much of the regulation is at posttranscriptional level (Whitfield et al., 2000) (Figure 1.1 (M. M. Koseoglu et al., 2010; Marzluff et al., 2008)). SLBP is required for all aspects of histone mRNA biosynthesis: proper processing of pre-mRNAs, nuclear export, translation and regulation of stability of mRNA (Dominski and Marzluff, 1999; Erkmann, Sanchez, et al., 2005; Gorgoni et al., 2005; Kaygun and Marzluff, 2005b; Sanchez and Marzluff, 2002; K. D. Sullivan et al., 2009; Whitfield et al., 2004). Mutation in stem loop results in no expression of processed histone mRNA in vivo (Ingledue et al., 1995; Levine et al., 1987, Pandey et al., 1994; Yang et al., 2009) most probably via abolishing SLBP binding. To date, there is no evidence for functions of SLBP outside those directly involved in histone biosynthesis (Townley-Tilson et al., 2006). Analysis of genetic mutations or knockdown of SLBP in Drosophila melanogaster (E. Sullivan et al., 2001), mouse (Arnold et al., 2008) and human cultured cells (Zhao et al., 2004) revealed that SLBP is required for cell cycle coupled histone mRNA production and indicate that the role of SLBP in histone production is highly conserved among different species. Other than in eukaryotes, SLBP was shown to be present in several types of protozoa (Davila Lopez and Samuelsson, 2008). Depletion of SLBP level in continuously cycling cells causes reduction in the rate of cell division, accumulation of cells in S phase probably via preventing sufficient histone production (K. D. Sullivan et al., 2009; Wagner et al., 2005; Zhao et al., 2004). One of the major defects caused in absence of SLBP is increase in the rate of formation of polyadenylated histone mRNAs, supporting the notion that SLBP is required for proper pre-mRNA processing (E. Sullivan et al., 2001). Expression of polyadenylated histone mRNAs have been revealed by microarray analyses in various subtypes of breast cancer (Zhao et al., 2004) and oropharyngeal squamous-cell carcinoma (Beggs et al., 2012; Feng et al., 2005; Kari et al., 2013). It is likely that, polyadenylated histone mRNAs might provide advantage to rapidly dividing cells by supplying additional source of histones outside of S phase, as they own cell cycle independent stability and translation (Kirsh et al., 1989).
One should expect that, restriction of SLBP expression just to S phase might require involvement of multiple regulatory mechanisms, as it is the core player of cell cycle regulation of histone biosynthesis. It was shown that rapid degradation of SLBP at S/G2 border is triggered due on phosphorylation by Cyclin A/CDK1 on Thr 61 which subsequently stimulates phosphorylation of adjacent Thr 60 by CK2 (M. M. Koseoglu et al., 2008). In our previous article (Djakbarova et al., 2016), we have identified the E3 Ligase, the CRL4DCAF11, which mediates SLBP degradation at S/G2 border, depending on double phosphorylation of SLBP. In order to identify the E3 Ligase, we have used 55 amino acid long SLBP piece, which is sufficient to mimic S/G2 regulation of SLBP expression (M. M. Koseoglu et al., 2008). By using this piece as a bait, we have found that CRL4DCAF11 binds to SLBP in phosphorylation dependent manner and mediates its degradation.
Followed by the rapid decline at the end of S phase, SLBP expression is repressed until next onset of S phase. Zheng et al, suggested that low translation efficiency was responsible for low SLBP level in G1 (Zheng et al., 2003), however it was shown that the low translation rate is limited to early G1 phase and it reaches to S phase level somewhere in between early to mid G1 ((Mehmet Murat Koseoglu, 2007) ; (Djakbarova et al., 2014). Unexpectedly, even after translation rate was enhanced, the level of SLBP does not reach to S phase level for couple of more hours, indicating the presence of additional mechanism with contrary effect. Previously proteasome mediated degradation was proposed that regulated proteasome mediated degradation is responsible for suppressed SLBP expression during G1 (Mehmet Murat Koseoglu, 2007). In our work, we have confirmed the proteasome mediated degradation suppresses SLBP expression low in G1 and showed that this degradation is independent from previously identified degradation at the end of S phase (Djakbarova et al., 2013).
Conclusıon.
SLBP is the major factor that regulates all aspects of histone mRNA metabolism. Restriction of SLBP expression to the S phase is a major mechanism to limit canonical histone production to the S phase. SLBP is rapidly degraded at the end of S phase as a mechanism to shut down canonical histone production.
REFERENCES
- Arias, E. E. and Walter, J. C., “Replication-dependent destruction of Cdt1 limits DNA replication to a single round per cell cycle in Xenopus egg extracts” // Genes Dev. - 2005. - Vol.19, №1. - Р. 114-126.
- Arias, E. E. and Walter, J. C., “PCNA functions as a molecular platform to trigger Cdt1 destruction and prevent re-replication” // Nat Cell Biol. - 2006. - Vol. 8, №1. - Р. 84-90.
- Arnold, D. R., Francon, P., Zhang, J., Martin, K. and Clarke, H. J., “Stem-loop binding protein expressed in growing oocytes is required for accumulation of mRNAs encoding histones H3 and H4 and for early embryonic development in the mouse” // Dev Biol. - 2008. - Vol. 313.№1. - Р. 347-358.
- Au, W. C., Crisp, M. J., DeLuca, S. Z., Rando, O. J. and Basrai, M. A., “Altered dosage and mislocalization of histone H3 and Cse4p lead to chromosome loss in Saccharomyces cerevisiae” // Genetics. - 2008. - Vol. 179, №1. - Р. 263-275.
- Avgousti, D. C., Palani, S., Sherman, Y. and Grishok, A., “CSR-1 RNAi pathway positively regulates histone expression in C. elegans” // Embo j. - 2012. - Vol. 31, №19. - Р. 3821-3832.
- Beggs, S., James, T. C. and Bond, U., “The PolyA tail length of yeast histone mRNAs varies during the cell cycle and is influenced by Sen1p and Rrp6p” // Nucleic Acids Res. - 2012. - Vol. 40, №6. - Р. 2700-2711.
- Busslinger, M., Portmann, R. and Birnsteil, M. L., “A regulatory sequence near the 3' end of sea urchin histone genes” // Nucleic Acids Res. Vol. 6, №9. - Р. 2997-3008.
- Cakmakci, N. G., Lerner, R. S., Wagner, E. J., Zheng, L. and Marzluff, W. F., “SLIP1, a factor required for activation of histone mRNA translation by the stem-loop binding protein” // Mol Cell Biol. - 2008. - Vol. 28, №3. - Р. 1182-1194.
- Connor, W., States, J. C., Mezquita, J. and Dixon, G. H., “Organization and nucleotide sequence of rainbow trout histone H2A and H3 genes” // J Mol Evol. - 1984. - Vol.20, №3(4). - Р. 236-250.
- Davila Lopez, M. and Samuelsson, T., “Early evolution of histone mRNA 3' end processing” // Rna. - 2008. - Vol. 14, №1. - Р. 1-10.
- DeJong, E. S., Marzluff, W. F. and Nikonowicz, E. P., “NMR structure and dynamics of the RNA-binding site for the histone mRNA stem-loop binding protein” // Rna. - 2002. - Vol. 8, №1. - Р. 83-96.
- DeLisle, A. J., Graves, R. A., Marzluff, W. F. and Johnson, L. F., “Regulation of histone mRNA production and stability in serum-stimulated mouse 3T6 fibroblasts” // Mol Cell Biol. - 1983. - Vol. 3, №11. - Р.1920-1929.
- Davila Lopez, M. and Samuelsson, T., “Early evolution of histone mRNA 3' end processing” // Rna. - 2008. - Vol. 14, №1. - Р. 1-10.
- DeJong, E. S., Marzluff, W. F. and Nikonowicz, E. P., “NMR structure and dynamics of the RNA-binding site for the histone mRNA stem-loop binding protein” // Rna. - 2002. - Vol. 8, №1. - Р. 83-96.
- DeLisle, A. J., Graves, R. A., Marzluff, W. F. and Johnson, L. F., “Regulation of histone mRNA production and stability in serum-stimulated mouse 3T6 fibroblasts” // Mol Cell Biol. - 1983. - Vol. 3, №11. - Р. 1920-1929.
- Djakbarova, U., Marzluff, W. F. and Koseoglu, M. M., “Translation regulation and proteasome mediated degradation cooperate to keep stem-loop binding protein low in G1-phase” // J Cell Biochem. - 2014. - Vol. 115, №3. - Р. 523-530.
- Djakbarova, U., Marzluff, W. F. and Koseoglu, M. M., “DDB1 and CUL4 associated factor 11 (DCAF11) mediates degradation of Stem-loop binding protein at the end of S phase” // Cell Cycle. - 2016. - Р. 1-11.
- Dollard, C., Ricupero-Hovasse, S. L., Natsoulis, G., Boeke, J. D. and Winston, F., “SPT10 and SPT21 are required for transcription of particular histone genes in Saccharomyces cerevisiae” // Mol Cell Biol. - 1994. - Vol. 14, №8. - Р. 5223-5228.
- Dominski, Z. and Marzluff, W. F., “Formation of the 3' end of histone mRNA” // Gene. - 1999. - Vol. 239, №1. - Р. 1-14.
- Dominski, Z. and Marzluff, W. F., “Formation of the 3' end of histone mRNA: getting closer to the end” // Gene. - 2007. - Vol. 396, №2. - Р. 373-390.
- Dominski, Z., Yang, X. C. and Marzluff, W. F., “The polyadenylation factor CPSF-73 is involved in histone-pre-mRNA processing” // Cell. - 2005. - Vol. 123, №1. - Р. 37-48.
- Erkmann, J. A., Sanchez, R., Treichel, N., Marzluff, W. F. and Kutay, U., “Nuclear export of metazoan replication-dependent histone mRNAs is dependent on RNA length and is mediated by TAP” // Rna. - 2005. - Vol. 11, №1. - Р. 45-58.
- Erkmann, J. A., Wagner, E. J., Dong, J., Zhang, Y., Kutay, U. and Marzluff, W. F., “Nuclear import of the stem-loop binding protein and localization during the cell cycle” // Mol Biol Cell. - 2005. - Vol. 16, №6. - Р. 2960-2971.
- Gallie, D. R., Lewis, N. J. and Marzluff, W. F., “The histone 3'-terminal stem-loop is necessary for translation in Chinese hamster ovary cells” // Nucleic Acids Res. - 1996. - Vol. 24, №10. - Р. 1954-1962.
- Ghule, P. N., Xie, R. L., Medina, R., Colby, J. L., Jones, S. N., Lian, J. B., Stein, J. L., van Wijnen, A. J. and Stein, G. S., “Fidelity of histone gene regulation is obligatory for genome replication and stability” // Mol Cell Biol. - 2014. - Vol. 34, №14. - Р. 2650-2659.
- Gick, O., Kramer, A., Keller, W. and Birnstiel, M. L., “Generation of histone mRNA 3' ends by endonucleolytic cleavage of the pre-mRNA in a snRNP-dependent in vitro reaction” // Embo j. - 1986. - Vol. 5, №6. - Р. 1319-1326.
- Graves, R. A., Pandey, N. B., Chodchoy, N. and Marzluff, W. F., “Translation is required for regulation of histone mRNA degradation” // Cell. 1987. - Vol. 48, №4. - Р. 615-626.
- Gunjan, A., Paik, J. and Verreault, A., “The emergence of regulated histone proteolysis” // Curr Opin Genet Dev. - 2006. - Vol. 16, №2. - Р. 112-118.
- Kaygun, H. and Marzluff, W. F., “Regulated degradation of replication-dependent histone mRNAs requires both ATR and Upf1” // Nat Struct Mol Biol. - 2005. - Vol. 12, №9. - Р. 794-800.
- Kaygun, H. and Marzluff, W. F., “Translation termination is involved in histone mRNA degradation when DNA replication is inhibited” // Mol Cell Biol. - Vol. 25, №16. - Р. 6879-6888.
- Kiriyama, M., Kobayashi, Y., Saito, M., Ishikawa, F. and Yonehara, S., “Interaction of FLASH with arsenite resistance protein 2 is involved in cell cycle progression at S phase” // Mol Cell Biol. - Vol. 29, №17. - Р. 4729-4741.
- Kirsh, A. L., Groudine, M. and Challoner, P. B., “Polyadenylation and U7 snRNP-mediated cleavage: alternative modes of RNA 3' processing in two avian histone H1 genes” // Genes Dev. - 1989. - Vol. 3, №12b. - Р. 2172-2179.
- Kohn, M. and Huttelmaier, S., “Non-coding RNAs, the cutting edge of histone messages” // RNA Biol. - 2016. - Vol. 13, №4. - Р. 367-372.
- Koseoglu, M. M., Cell cycle regulation of Stem-Loop Binding Protein: a key regulator in histone mRNA metabolism. , Ph.D dissertation, University of North Carolina at Chapel Hill. - 2007. - 108 р.
- Koseoglu, M. M., Dong, J. and Marzluff, W. F., “Coordinate regulation of histone mRNA metabolism and DNA replication: cyclin A/cdk1 is involved in inactivation of histone mRNA metabolism and DNA replication at the end of S phase” // Cell Cycle. - 2010. - Vol. 9, №19. - Р. 3857-3863.
- Koseoglu, M. M., Graves, L. M. and Marzluff, W. F., “Phosphorylation of threonine 61 by cyclin a/Cdk1 triggers degradation of stem-loop binding protein at the end of S phase” // Mol Cell Biol. - 2008. - Vol. 28, №14. - Р. 4469-4479.
- Levine, B. J., Chodchoy, N., Marzluff, W. F. and Skoultchi, A. I., “Coupling of replication type histone mRNA levels to DNA synthesis requires the stem-loop sequence at the 3' end of the mRNA” // Proc Natl Acad Sci U S A. - 1987. - Vol. 84, №17. - Р. 6189-6193.
- Marzluff, W. F., “Terminating histone synthesis to preserve centromere integrity” // Dev Cell. - 2010. - Vol. 18, №3. - Р. 335-336.
- Marzluff, W. F. and Duronio, R. J., “Histone mRNA expression: multiple levels of cell cycle regulation and important developmental consequences” // Curr Opin Cell Biol. - Vol. 14, №6. - Р. 692-699.
- Marzluff, W. F., Gongidi, P., Woods, K. R., Jin, J. and Maltais, L. J., “The human and mouse replication-dependent histone genes” // Genomics. - Vol. 80, №5. - Р. 487-498.
- Raczynska, K. D., Ruepp, M. D., Brzek, A., Reber, S., Romeo, V., Rindlisbacher, B., Heller, M., Szweykowska-Kulinska, Z., Jarmolowski, A. and Schumperli, D., “FUS/TLS contributes to replication-dependent histone gene expression by interaction with U7 snRNPs and histonespecific transcription factors” // Nucleic Acids Res. - 2015. - Vol. 43, №20. - Р. 9711-9728.
- Sullivan, K. D., Mullen, T. E., Marzluff, W. F. and Wagner, E. J., “Knockdown of SLBP results in nuclear retention of histone mRNA” // Rna. - 2009. - Vol. 15, №3. - Р. 459-472.
- Wagner, E. J., Berkow, A. and Marzluff, W. F., “Expression of an RNAi-resistant SLBP restores proper S-phase progression” // Biochem Soc Trans. - 2005. - Vol. 33, №3. - Р. 471-473.
- Whitfield, M. L., Kaygun, H., Erkmann, J. A., Townley-Tilson, W. H., Dominski, Z. and Marzluff, W. F., “SLBP is associated with histone mRNA on polyribosomes as a component of the histone mRNP” // Nucleic Acids Res. - 2004. - Vol. 32, №16. - Р. 4833-4842.
- Whitfield, M. L., Zheng, L. X., Baldwin, A., Ohta, T., Hurt, M. M. and Marzluff, W. F., “Stem-loop binding protein, the protein that binds the 3' end of histone mRNA, is cell cycle regulated by both translational and posttranslational mechanisms” // Mol Cell Biol. - Vol. 20, №12. - Р. 4188-4198.
- Yang, X. C., Torres, M. P., Marzluff, W. F. and Dominski, Z., “Three proteins of the U7-specific Sm ring function as the molecular ruler to determine the site of 3'-end processing in mammalian histone pre-mRNA” // Mol Cell Biol. - 2009. - Vol. 29, №15. - Р. 4045-4056.
- Zhang, J., Tan, D., DeRose, E. F., Perera, L., Dominski, Z., Marzluff, W. F., Tong, L. and Hall, T. M., “Molecular mechanisms for the regulation of histone mRNA stem-loop-binding protein by phosphorylation” // Proc Natl Acad Sci U S A. - 2014. - Vol. 111, №29. - Р. 2937-2946.
- Zhang, M., Lam, T. T., Tonelli, M., Marzluff, W. F. and Thapar, R., “Interaction of the histone mRNA hairpin with stem-loop binding protein (SLBP) and regulation of the SLBP-RNA complex by phosphorylation and proline isomerization” // Biochemistry. - 2012. - Vol. 51, №15. Р. 3215-3231.
- Zhao, X., McKillop-Smith, S. and Muller, B., “The human histone gene expression regulator HBP/SLBP is required for histone and DNA synthesis, cell cycle progression and cell proliferation in mitotic cells” // J Cell Sci. - 2004. - Vol. 117, №25. - Р. 6043-6051.
- Zheng, L., Dominski, Z., Yang, X. C., Elms, P., Raska, C. S., Borchers, C. H. and Marzluff, W. F., “Phosphorylation of stem-loop binding protein (SLBP) on two threonines triggers degradation of SLBP, the sole cell cycle-regulated factor required for regulation of histone mRNA processing, at the end of S phase” // Mol Cell Biol. - 2003. - Vol. 23, №5. - Р. 1590-1601.