The goal of modern medicine is early diagnosis socially significant diseases that reduce the ability to work and affect on mortality in the population. Oncological diseases occupy second rank among the causes of deaths in Kazakhstan. At the same time, lung cancer is leading, the death rate from which in the Republic is 17.6 % of the total mortality from cancer. The cause of high mortality is the lack of effective diagnosis of lung cancer in the early stages of the disease. At the moment, clinical medicine does not have specific and highly sensitive oncomarkers suitable for using in screening studies of the population. Free-circulating nucleic acids, including mitochondrial DNA and microRNA, are stable in physiological fluids of the body, such as blood, serum, saliva, urine, etc. Moreover, the results of numerous studies have shown that the level of free- circulating nucleic acids can indicate pathological processes in the human body. This shows great prospects for the use of free-circulating nucleic acids as biomarkers for non-invasive diagnosis of cancer. In this review, we are presenting available data on the modern methods of non-invasive cancer detection, so called «liquid biopsy» based on free circulating nucleic acids.
Currently, one of the problems of modern medicine is the growth of cancer rate in the developed countries, including Kazakhstan. Among malignant diseases the lung cancer occupies a leading position [1].
The lung cancer in the Republic of Kazakhstan makes up 11.4 % of the total number of malignant tumors. By morbidity and mortality, lung cancer ranks first place in Kazakhstan [2]. Exposure to radon, a naturally existing radioactive gas, is the second leading cause, according to the World Health Organization [3]. According to the experts the contribution of natural sources in the average annual radiation dose of the Kazakh population currently stands at 80 %, including 50 % from radon [4]. The genotoxic effects of radon on population of Kazakhstan are poorly understood, in spite of the fact that many regions of the country contain the high levels of radon. Therefore the search for blood-based tumor biomarkers for early detection of lung cancer is particularly acute in Kazakhstan.
Liquid biopsy is a revolutionary technique that is opening previously unexpected perspectives. It consists of the detection and isolation of free-circulating DNA, as a source of genomic information in patients with cancer.
In addition to the widely used methods of clinical diagnostics of cancer diseases, to date, a large amount of evidence has been accumulated about the involvement of microRNA in the carcinogenesis of various tumors, including lung cancer [5].
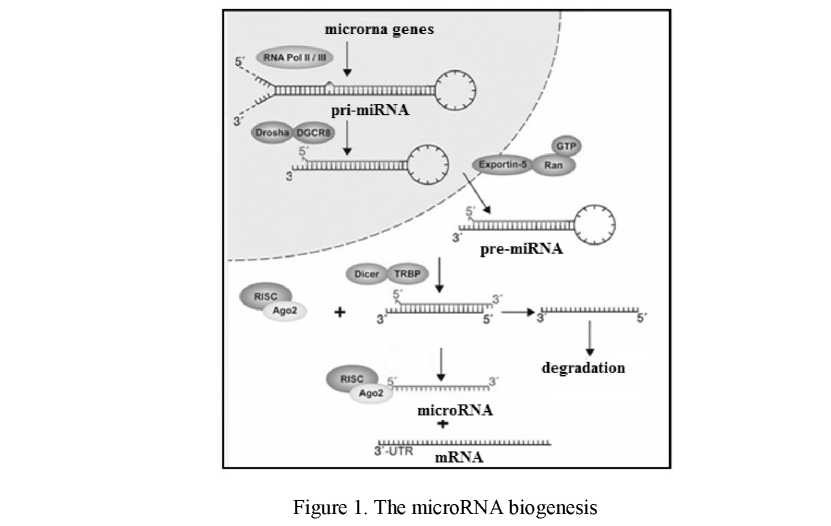
MicroRNAs (miRNAs) are small (~22 nucleotides), non-coding RNA molecules that regulate gene expression post-transcriptionally by inhibiting target mRNAs (Fig. 1).
As a result of the transcription of microRNA' genes, pri-microRNA is formed, having a length of a thousand base pairs. The nucleotide sequence of these molecules includes inverted repeats, due to which a hairpin structure forms [6].
The microprocessor protein complex consisting of proteins are known as Drosha and Pasha. Pasha or its analog in mammalian cells DGCR8, recognizes the features of the pri-microRNA' structure. In the opinion of some authors, Pasha/DGCR8 recognizes the cleavage sites of pri-microRNA due to the presence in their structures an extended terminal loop [7]. According to other authors, site identification is associated with the transition point of the double-stranded RNA structure to flanking single-stranded areas [8] (Fig. 1). DROSHA cleaves the 3' and 5' strands of a stem-loop in pri-miRNAs. After the events described above, occurring in the nucleus, pre-microRNA is formed. Pre-microRNA has two free nucleotides at the 3'-end, which are necessary for subsequent interaction with the Exportin 5 is a RanGTP-dependent protein that mediates nuclear export of pre-miRNAs.
Moreover the free nucleotides at the 3'-end are necessary for the interaction of pre-microRNA with a protein called Dicer. This enzyme, which contains the catalytic domain of RNase III, cuts out of the pre- microRNA' loop that connects the 3'- and 5'-ends of the hairpin. As a result, short double-stranded RNA molecules are formed [9].
The last stage of biogenesis of microRNA is mediated by the conversion of double-stranded RNA to mature single-stranded RNA (Fig. 1), which is used to recognize their mRNA targets. This process takes place with the participation of the RISC complex (RNA Induced Silencing Complex), which includes one of the proteins of the Argonaute family (AGO1, AGO2). microRNA-RISC complex leads to degradation of mRNA or block translational initiation [10].
An estimated one third of the human genome is regulated by microRNAs. Differentially expressed mi- croRNAs have been identified in bronchopulmonary dysplasia, many of which regulate late lung development and alveolarization. Evidence is increasing that microRNAs are particularly important in lung homeostasis and development and have been demonstrated to be involved in many pulmonary diseases such as astma, COPD, lung cancer and other smoking related diseases.
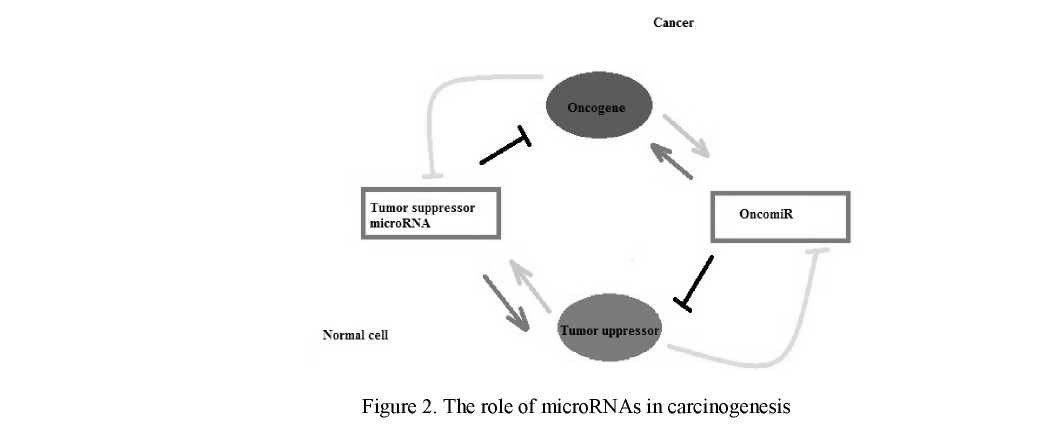
The important role of a differential expression of genes of microRNA in processes of malignant transformation is shown, and profiles of an expression of microRNA carry a tumor — specific character [11]. Recently, some microRNA-based drugs have been introduced as possible therapeutics agents. microRNAs act as a converter / mediator in the gene network of oncogenes and onco-suppressors. In one case, microRNA, regulated by oncogenes, can turn off onco- suppressor genes, which allows weakening cellular control over proliferation and adhesion. In another case, the tumor-suppressor microRNA can suppress the action of the oncogene (Fig. 2). Recently, some microRNA-based drugs have been introduced as possible therapeutics agents.
We have studied the level of a microRNAs expression in lung cancer patients in comparison with healthy people [5, 12, 13]. The comparative analysis of blood free circulating microRNA in group of lung cancer patients vs control group showed the high level of some oncomiRs such as miR-155-5р [12], miR-19b-3p [5] and miR-125b-5p [13]. Our results demonstrate that expression of this microRNA in lung cancer patients was in 2, 7 and 4 times higher respectively than in control group. That allows to draw a conclusion about the possibility of using free-circulating microRNA as biomarkers of lung cancer [11, 14].
The interesting data were obtained about the effect of smoking as the main risk factor for the development of lung cancer and on the level of oncomiRs expression [5, 15]. In our studies the obtained results demonstrated, that in the smoking group of lung cancer patients the expression level of miR-19b-3p and miR-155 -5p was significantly lower than in the group of non-smokers lung cancer patients. It is possible that this effect is explained a general decrease in all freely circulating miRNAs due to the inactivation of DICER, which is the main enzyme of miRNA biogenesis, by tobacco smoke. Our results are consistent with previous data, indicating the negative effect of tobacco smoke on miRNA biogenesis [16, 17].
Emerging evidence is establishing that microRNA are involved in regulating radiation-induced cellular processes and can be exploited to serve as biomarkers of human radiation exposure [18]. Recently, it has been shown that ionizing radiation can induce changes in miRNA expression profiles in normal human fibroblasts [18] and immortalized cell lines [19, 20]. An in vivo study found that microRNA signatures induced by ionizing radiation in mouse blood are radiation type and dose-specific [21]. microRNA expression signatures derived from mouse blood are exploited to serve as biomarkers for exposure to radiation [20]. Another study conducted on radiotherapy patients suggested that microRNA expression signatures can be used as biomarkers of radiation exposure in humans [21].
MicroRNAs are very promising as cancer biomarkers, but additional studies, especially meta-analyses of pooled data from different studies, are needed to validate the efficacy of microRNAs and to identify reliable microRNA panels to be used as cancer biomarkers.
The results of recent studies show that the free circulating mitochondrial DNA (mtDNA) also can be used as one of the promising markers for pathological processes' detection. At the present time it has become apparent that mitochondrial dysfunctions determine the development of many diseases, including malignant tumors and autoimmune diseases [22, 23].
Mitochondrial dysfunction leads to production and accumulation of the reactive oxygen species (ROS) which effect on the processes in the cells [24]. The accumulation of ROS in cells leads to oxidative stress and can break the work of the mitochondrial enzyme systems. In normal physiology, mitochondria are very important in the cell as they produce most of the adenosine triphosphate (ATP) via oxidative phosphorylation system which is a necessary energy supply for cellular processes.
It is known that in the hypoxia conditions normal functions of the mitochondrial respiratory chain complexes is disturbed, which is accompanied by increasing of mitochondrial membrane permeability. This process is responsible for the exit of mitochondrial DNA into the cytosol.
It is shown that in cancer cells increasing an anaerobic glycolysis is observed. Also the number of mitochondrial DNA copies changes at various forms of cancer. The increased number of copies mtDNA was found in a prostate and ovaries cancer, endometrial cancer and some forms of leukemia. To the contrary, the reduced number of copies mtDNA was found in gastric and liver carcinoma, in a breast cancer and other forms of cancer [25].
Mutations in the D-loop region were found both in mtDNA, directly isolated from liver carcinoma cells, and in the free-circulating mtDNA in the blood plasma of patients with liver cancer, which indicates that free-circulating mtDNA exists in the common free circulating DNA pool [26]. Whereas circulating mtDNA was also found in the blood of healthy people, as well as in patients with various diseases [27], particularly significant differences in the level of free circulating mtDNA were found between cancer patients and healthy individuals [10]. This made it possible to use the quantitative analysis of the mtDNA copy number in biological fluids (plasma / blood / urine) for early detection of different types of cancer [28].
Subsequent studies have shown that the level of free-circulating mtDNA can be used as a marker for the survival prognosis of patients with malignant tumors of different locations. Thus, a high level of free circulating mtDNA correlated with low survival of patients with prostate cancer [29].
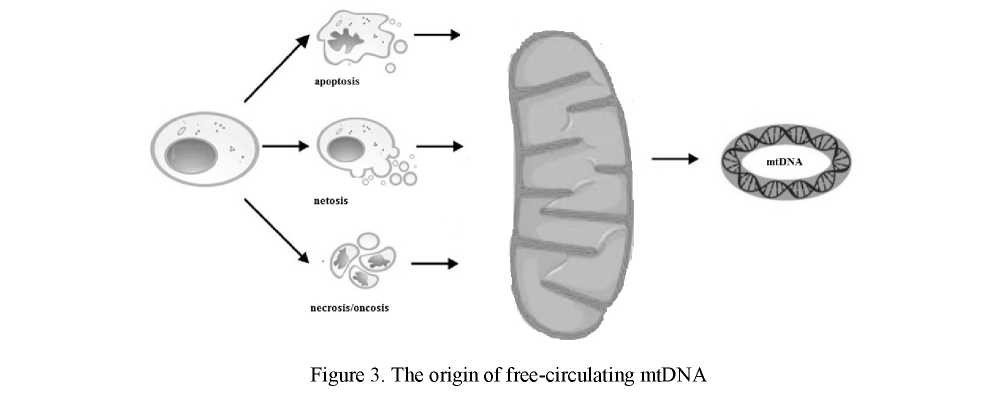
Mitochondrial DNA is released in the circulation following general mechanisms of cell death: apoptosis and necrosis (Fig.3). Oncosis is a non-canonical form of programmed cell also can cause the presence of free-circulating mtDNA in the blood plasma of cancer patients (Fig. 3) [30].
Given that under certain conditions, neutrophils form mtDNA neutrophil extracellular traps (NETs), it is feasible that the circulating mtDNA observed may have been released through netosis (Fig. 3).
The recent data suggest that mtDNA can also induce the non-infectious systemic inflammatory response [30]. As we know that inflammation is a critical component of tumour progression. Therefore, it is quite possible that mtDNA can be not only a consequence, but also a cause of carcinogenesis.
Free-circulating mtDNA can be an indicator of carcinogenic effects [31]. Tan and colleagues show that cigarette smoke leads to increasing the frequency of mutations in the coding regions and in the D-loop region of mtDNA in buccal cells of smokers compared to the non-smokers [32]. Moreover, tobacco smoking causes an increase in the level of free-circulating mtDNA in the plasma of patients with malignant tumors of the head and neck [33].
The results of several studies indicate the possible role of circulating mtDNA as a biomarker of cellular damage caused by chronic exposure to low radiation doses [34]. Animal studies showed a significant increasing level of free-circulating mtDNA after X-ray exposure. In addition, the results of this experiment detected a number of mutations in free-circulating mtDNA after X-ray irradiation, which were absent in the control group without X-ray exposure [35].
The results of several studies indicate the possible role of circulating mtDNA as a biomarker of cellular damage caused by chronic exposure to low radiation doses [33]. Animal studies showed a significant increasing level of free-circulating mtDNA after X-ray exposure. In addition, the results of this experiment detected a number of mutations in free-circulating mtDNA after X-ray irradiation, which were absent in the control group without X-ray exposure [35].
Radiation therapy of cancer diseases also leads to increasing the level of free-circulating mtDNA in the plasma of patients and, consequently, this indicator can be used to monitor the effectiveness of treatment with ionizing radiation of cancer patients [36].
However, the influence of environmental factors on the level of free-circulating mtDNA at the moment is still not understood. Further research in this area can contribute to the development of preventive diagnostics aimed at identifying potential health risks associated with the impact of adverse environmental conditions or an incorrect lifestyle.
Several studies have shown that the level of the free-circulating mtDNA can be use as a biomarker of the cancer therapeutic resistance. For example, two independent studies have shown that a decrease in the level of free-circulating mtDNA associated with resistance to antitumor drugs such as anthracyclines and taxanes [37, 38]. Moreover Sanson et al., have shown that exogenous mtDNA can act as an oncogenic signal that promotes the activation of cancer stem cells after therapy. The horizontal transfer of mtDNA in the form of extracellular vesicles in contrast, provoked the development of resistance to hormonal therapy for breast cancer [39].
The correlation between the low level of mtDNA in the patient's plasma and the low susceptibility of the tumor to the drugs can be explained by the absence of death of tumor cells, which mediates the massive yield of free-circulating mtDNA, i.e. cancer therapy does not induce apoptosis / necrosis of tumor cells. Consequently, free-circulating mtDNA as a kind of marker of cell death can be used to monitor the effectiveness of the cancer therapy. A decrease in the level of free-circulating mtDNA in the blood plasma of lung cancer patients was shown in the case of resistance to erlotinib [40].
For many years, researchers have tried to create effective methods for cancer diagnosis, allowing detected malignant tumor at an early stage of development. Moreover, these diagnostic methods should be characterized by high specificity, sensitivity, absence of complications and to be sufficiently simple in technical terms. At the moment, molecular technology called «liquid biopsy» is becoming increasingly popular. A tissue biopsy is the «gold standard» of modern oncology. Despite the high accuracy, this method is very expensive, traumatic and unacceptable for mass screening studies of the population. «Liquid biopsy» is based on the use of physiological body fluids, such as blood, urine, saliva, etc., and has no risks for the patient compared to tissue biopsy. The markers in this case are free-circulating nucleic acids: microRNA and mitochondrial DNA.
Diagnostic methods based on the detection of free-circulating nucleic acids can be used not only in screening studies, but also serve as a reliable marker for the prognosis of the disease and the effectiveness of cancer therapy.
Thus, understanding the nature of these molecules, the mechanism of the biogenesis of free-circulating nucleic acids and their involvement in the processes of carcinogenesis is importance for personalized treatment of cancer diseases.
References
- Aggarwal, A., Lewison, G., Idir, S., Peters, M., Aldige, C., & Boerckel, W. (2016). The state of lung cancer research: a global analysis. J. Thorac. Oncol., 11, 10-21.
- Bersimbaev, R., & Bulgakova, O. (2017). Residential radon exposure and lung cancer risk in Kazakhstan. Radon. F. Adrovic (ed.). InTech, Rijeka, Croatia, 93-124.
- Zeeb, H., & Shannon, F. (2009). WHO Handbook on Indoor Radon: A Public Health Perspective. World Health Organization. Zeeb H and Shannon F (eds.). World Health Organization, Geneva, 3-21.
- Bersimbaev, R., & Bulgakova, O. (2015). The health effects of radon and uranium on the population of Kazakhstan. Genes and Environment 37, 18. doi: 10.1186/s41021-015-0019-3.
- Bulgakova, O., Zhabayeva, D., Kussainova, A., Pulliero, A., Izzotti, A., & Bersimbaev, R. (2018). miR-19 in blood plasma reflects lung cancer occurrence but is not specifically associated with radon exposure. Oncology Letters, 15(6), 8816-8824.
- Wahid, F., Shehzad, A., Khan, T., & Kim, Y.Y. (2010). MicroRNAs: synthesis, mechanism, function, and recent clinical trials. BiochimBiophys Acta, 1803(11), 1231-1243.
- Zeng, Y., Yi, R., & Cullen, B.R. (2005). Recognition and cleavage of primary microRNA precursors by the nuclear processing enzyme Drosha. EMBO J., 24(1), 138-148.
- Han J., Lee Y., Yeom K. H., Nam J. W., & Heo I. (2006). Molecular basis for the recognition of primary microRNAs by the Drosha-DGCR8 complex. Cell, 125(5), 887-901.
- Chendrimada, T.P. et al. (2005). TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature, 436(7051), 740-744.
- Janowski, B.A. et al. (2006). Involvement of AGO1 and AGO2 in mammalian transcriptional silencing. Nat. Struct. Mol. Biol., 13(9), 787-792.
- Izotti, A., Carozzo, S., Pulhiero, A., Zhabayeva, D., Ravetti, J.L., & Bersimbaev, R.I. (2016). Extracellular microRNA in liquid biopsy: applicability in cancer diagnosis and prevention. Amer. J. Cancer Res., 6(7), 1461-1493.
- Bulgakova, O.V., Zhabaeva, D.B., & Bersimbayev, R.I. (2017). Rol miR-155-5p v patoheneze raka lehkoho [Role of miR- 155-5p in the pathogenesis of lung cancer]. Doklady NAN RK — Reports of NAS RK, 3, 121-129 [in Russian].
- Bulgakova, O.V., & Bersimbayev, R.I. (2017). Rol miR125b v patoheneze raka lehkoho [Role miR125b in the pathogenesis of lung cancer] Proceedings from Postgenomic technologies in medicine: from theory to practice'17. Konferentsiia — Conference (pp. 86-92). Voronezh [in Russian].
- Izotti, A., Geretto, M., Pulliero, A., Zhabayeva, D., & Bersimbaev, R.I. (2017). Resistance to cancer chemotherapeutic drugs is determined by pivotal microRNA regulators. Amer. J. Cancer Res., 7(7), 1275-1297.
- Bulgakova, O.V., & Bersimbaev, R.I. (2017). Analiz ekspressii mikroRNK pri rake lehkoho, indutsirovannoho kureniem [Analysis of microRNA expression in lung cancer induced by smoking]. Uspekhi molekuliarnoi onkolohii — Advances in molecular oncology, 4, 33–34 [in Russian].
- Gross, T., Powers, L., Boudreau, R., Brink, B., Reisetter, A., & Goel, K., et al. (2014). A MicroRNA Processing Defect in Smokers' Macrophages Is Linked to SUMOylation of the Endonuclease DIcER. J. Biol. Chem., 289(18), 12823–12834.
- Izzotti, A., Balansky, R., Ganchev, G., Iltcheva, M., Longobardi, M., & Pulliero, A., et al. (2016). Blood and lung mi- croRNAs as biomarkers of pulmonary tumorigenesis in cigarette smoke-exposed mice. Oncotarget, 7(51), 84758–84774.
- Simone, N.L., Soule, B.P., & Ly, D., et al. (2009). Ionizing radiation-induced oxidative stress alters microRNA expression. PLoS. One, 4, E6377.
- Shin, S., cha, H.J., & Lee, E.M., et al. (2009). Alteration of miRNA profiles by ionizing radiation in A549 human non-small cell lung cancer cells. Int. J. Oncol., 35, 81–86.
- chaudhry, M.A. (2009). Real-time PcR analysis of micro-RNA expression in ionizing radiation-treated cells. Cancer BiotherRadiopharm, 24, 49–56.
- Templin, T. Paul, S., Amundson, S.A., Young, E.F., Barker, c.A., Wolden, S.L., & Smilenov, L.B. (2011). Radiation- induced micro-RNA expression changes in peripheral blood cells of radiotherapy patients. Int. J. Radiat. Oncol. Biol. Phys., 80(2), 549–557.
- Holdenrieder, S. (2010). New challenges for laboratory diagnostics in non-small cell lung cancer. Cancer Biomarkers, 6, 3–4, 119–121.
- Frank, M.O. (2016). circulating cell-free DNA differentiates severity of inflammation. Biol. Res. Nurs. pii: 1099800416642571.
- Berezin, A.E. (2017). circulating cell-free mitochondrial DNA as biomarker of cardivascular risk: new challenges of old findings. Angiology, 3(4), 1–3.
- Meng, A. (2013). Mitochondrial DNA copy number alteration in human cancers. NA J. Med. Sci., 6(1), 22–25.
- Nomoto, S., Yamashita, K., Koshikawa, K., Nakao, A., & Sidransky, D. (2002). Mitochondrial D-loop mutations as clonal markers in multicentric hepatocellular carcinoma and plasma. Clin. Cancer Res., 8(2), 481–487.
- chiu, R.W.K., chan, L.Y.S., Lam, N.Y.L., Tsui, N.B.Y., Ng, E.K.O., & Rainer, T.H., et al. (2003). Quantitative analysis of circulating mitochondrial DNA in plasma. Clinical Chemistry, 4(5), 719–726.
- Yu, M. (2012). circulating cell-free mitochondrial DNA as a novel cancer biomarker: opportunities and challenges. Mitochondrial DNA, 23(5), 329–332.
- Mehra, N., Penning, M., Maas, J., van Daal, N., Giles, R.H., & Voest, E.E. (2007). circulating mitochondrial nucleic acids have prognostic value for survival in patients with advanced prostate cancer. Clin. Cancer Res., 13, 421–426.
- Bulgakova, O.V. (2018) Sovremennye predstavleniia o svobodno-tsirkuliruiushchikh nukleinovykh kislotakh i ikh rol v meditsine. [The contemporary views of free-circulating nucleic acids and their role in medicine]. Astana: Master Po, 114 [in Russian].
- Budnik, L.T., Kloth, S., Baur, X., Preisser, A.M., & Schwarzenbach, H. (2013). circulating mitochondrial DNA as biomarker linking environmental chemical exposure to early preclinical lesions elevation of mtDNA in human serum after exposure to carcinogenic halo-alkane-based pesticides. PloS. One, 8(5), E64413.
- Tan, D., Goerlitz, D.S., & Dumitrescu, R.G. et al. (2008). Associations between cigarette smoking and mitochondrial DNA abnormalities in buccal cells. Carcinogenesis, 29(6), 1170–1177.
- Kumar, M., Srivastava, S., Singh, S.A., Das, A.K., Das, G.c., & Dhar, B., et al. (2017). cell-free mitochondrial DNA copy number variation in head and neck squamous cell carcinoma: A study of non-invasive biomarker from Northeast India. Tumour Biol., 39(10), DOI: 1010428317736643.
- Borghini, A., Mercuri, A., Turchi, S., chiesa, M.R., Piccaluga, E., & Andreassi, M.G. (2015). Increased circulating cell-free DNA levels and mtDNA fragments in interventional cardiologists occupationally exposed to low levels of ionizing radiation. Environ. Mol. Mutagen, 56(3), 293–300.
- Abdullaev, S.A., Minkabirova, G.M., Bezlepkin, V.G., & Gaziev, A. (2015). cell-free DNA in the urine of rats exposed to ionizing radiation. Radiat. Environ. Biophys., 54(3), 297–304.
- cheng, c., Omura-Minamisawa, M., Kang, Y., Hara, T., Koike, I., & Inoue, T. (2009). Quantification of circulating cell-free DNA in the plasma of cancer patients during radiation therapy. Cancer Sci., 100(2), 303–309.
- Guerra, F., Perrone, A.M., Kurelac, I., Santini, D., ceccarelli, c., & cricca, M., et al. (2012). Mitochondrial DNA mutation in serous ovarian cancer: implications for mitochondria-coded genes in chemoresistance. Clin. Oncol., 30(36), E373-E378.
- Hsu, c.W., Yin, P.H., Lee, H.c., chi, c.W., & Tseng, L.M. (2010). Mitochondrial DNA content as a potential marker to predict response to anthracycline in breast cancer patients. Breast J., 16(3), 264-270.
- Sansone, P., Savini, c., Kurelac, I., chang, Q., Amato, L.B., & Strillacci, A., et al. (2017). Packaging and transfer of mitochondrial DNA via exosomes regulate escape from dormancy in hormonal therapy-resistant breast cancer. Proceedings of the National Academy of Sciences of the United States of America, 114(43), E9066-E9075.
- Huang, c-Y., chen, Y-M., & Wu, c-H. et al. (2014). circulating free mitochondrial DNA concentration and its association with erlotinib treatment in patients with adenocarcinoma of the lung. Oncology Letters, 7(6), 2180-2184.