DNA extraction is a crucial step at using molecular and biological methods, since the result of PCR analysis largely depends on the quality and quantity of the obtained DNA preparations. In this work, we compared the efficiency of DNA extraction from bacteria in various ways: the trizol method using TRizol commercial reagent, the CTAB method and the sorbent method using silica particles. It has been shown that the extraction of DNA from Salmonella bacteria on silica particles ensures higher sensitivity than the other methods. The amplification of the extracted DNA on silica particles is more efficient and allows obtaining reproducible results with the accuracy of a high degree. In the future, this method will be used to complete the test system for diagnosing salmonellosis. Introduction of the methods of express diagnosis into practice is extremely important, as quick and early diagnosis will allow counteracting the spread of infection in a timely manner, which in turn will improve the epidemiological and epizootological situation in the country.
Introduction
Salmonellosis is a widely spread human and animal infection caused by various members of the Salmonella genus. Food raw material and foodstuff are the most dangerous for people of Salmonella reservoir. In many European countries, salmonellosis morbidity among humans has significantly increased for two recent decades [1].
Lately, methods of gene diagnosis (nucleic acid hybridization, polymerase chain reaction — PCR, and so on) are successfully used to detect and identify microorganisms in various objects under study. These methods allow indicating and identifying microorganisms at high specificity in the presence of attendant microflora and with sensitivity right up to single cells [2].
The identification of infectious agents by classical bacteriological methods is associated with the usage of numerous selective nutritive media and various substrates. It makes the identification process protracted and laborious. Moreover, growing contaminating microorganisms in various media under competition conditions not always results in adequate identification. Furthermore, in vivo identification of transformed bacteria or of artificially genetically changed forms carrying pathogenicity genes not typical for the strain is difficult in the classical bacteriological assay [3].
Currently, there are various methods that allow extracting nucleic acids from a wide range of samples but only a bit of them is suitable for automation and many steps of isolation involve the risk of contamination. The presence of contaminants (proteins, for instance) in the complex mixtures often impedes the implementation of necessary reactions and techniques. Methods of DNA or RNA extraction should meet the following priority requirements: lysis of the biological material, selective extraction (sorption), concentration from large volumes, separation of components that inhibit PCR, separation of DNA and RNA, high yield percentage, opportunity of calibration and of positive control, absence of contamination, low time expenses, automation possibility [4].
Dynamic of development of veterinary and sanitary expert examination and of foodstuff certification is characterized by transfer to the most specific, sensitive and accelerated methods of bacteria identification [5].
Polymerase chain reaction (PCR) has a number of advantages as compared to the traditional bacteriological method of Salmonella detection since it combines rapidness and simplicity of implementation as well as potentially high specificity and sensitivity in revealing pathogenic microorganisms [6].
The objective of our studies was to select an effective method of DNA extraction for Salmonella detection by the method of polymerase chain reaction.
Materials and Methods
The objects of the study were eight Salmonella bacteria isolated from the foodstuff at the Microbiology Chair of Al-Farabi Kazakh National University.
DNA Extraction. DNA was extracted from Salmonella bacteria by the following methods:
- Trizol method. The commercial reagent TRizol («Invitrogen», USA) was used in accordance with the manufacturer's manual.
- CTAB method. CTAB (cetyl trimethyl ammonium bromide) is a classical cation detergent that is used in DNA extraction. СТАВ lyses the cellular membrane, effectively destroys DNA-protein [7].
- Sorbent method using silica particles. DNA binds to silica particles under the effect of high concentrated salt [8].
The quality and quantity of the extracted DNA were controlled with the use of NanoDrop 2000.
For setting PCR used the same amount of DNA (~ 30 ng/µɪ) obtained by different methods.
PCR-amplification. PCR-fragments of Inv gene from Salmonella bacteria were produced with the use of the following primers: S Inv-1F (direct) and S Inv-1R (inverse). Amplification was carried out in 25 µL of reaction mixture of the following composition: 10 xbuffer of DNA polymerase — 2.5 µL, 10 mM dNTP — 1 µL, MgCl2 — 1 µL, 20-50 ng of DNA-matrix, 20 pM of direct and 20 pM of inverse primers, and 0.5 U of DNA polymerase (Invitrogen), DNA — 1 µL. PCR products were assayed by electrophoresis in 1.5 % agarose gel containing ethidium bromide (1 µg/µL) at the field strength 6 V/cm2. The PCR product sized 500 bp.
Results and Discussion
Comparative study of three different versions of the DNA extraction from Salmonella bacteria was carried out: the trizol method using TRizol commercial reagent, the CTAB method and the sorbent method using silica particles.
The parameters such as quantity and quality of the obtained DNA were taken into consideration to select the DNA extraction method. The results of extracting genome DNA by different methods from 8 Salmonella isolates are shown in Figure 1.
The implemented studies showed the CTAB method to demonstrate the highest DNA yield. The results of the trizol method are slightly lower. And the lowest DNA yield was demonstrated by the sorbent method.
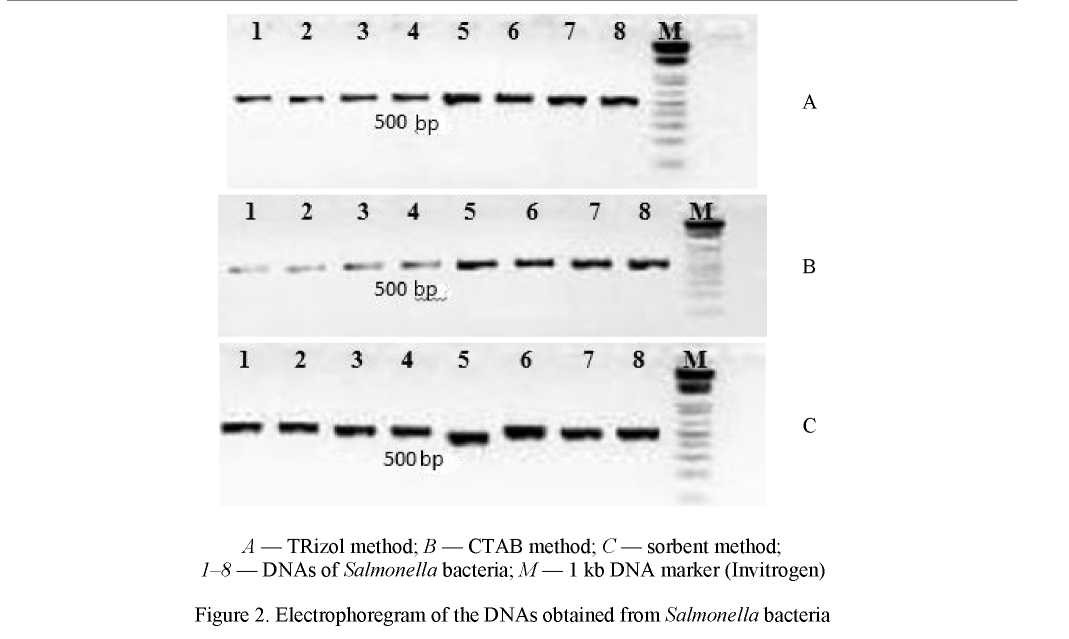
PCR-assay depends significantly on the quality of the extracted DNA. PCR amplification was conducted with the DNA specimens of equal volume (1 µL) obtained by different extraction methods from samples of the materials under study: the trizol method using TRizol commercial reagent, the CTAB method and the sorbent method using silica particles. The results are shown in Figure 2.
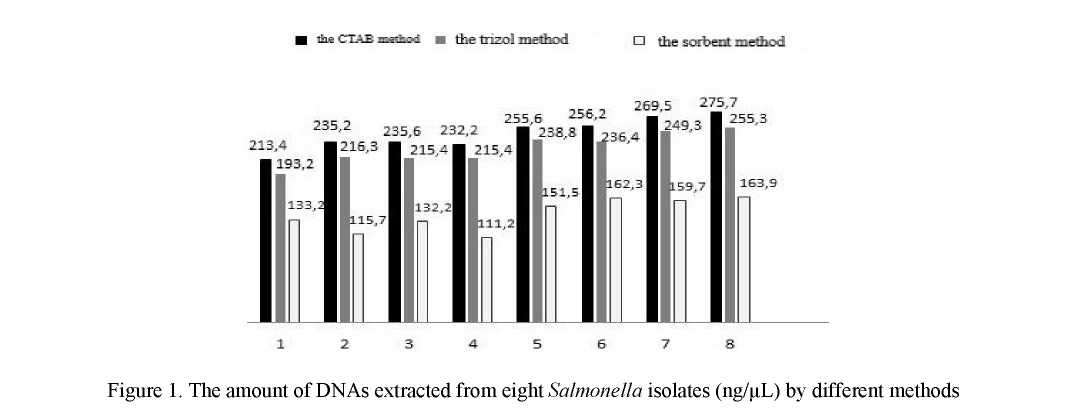
Figure 2 shows that the genetic material of high quality and of concentration sufficient for PCR-assay has been obtained by the sorbent method with the use of silica particles (С). At the same time, it should be noted that the DNA concentration in these preparations is 1.61 times less than in preparations obtained by using TRizol (A) and 1.75 times lower versus the preparations obtained by the CTAB method.
Insignificant PCR inhibition in the DNA specimens obtained by the trizol method using TRizol commercial reagent and the CTAB method may be caused by the DNA excess, which is a PCR inhibitor.
The same amount of DNA (50 ng/µŋ was used to select an effective DNA extraction method for detecting Salmonella in the PCR reaction. The results are presented in Figure 3.
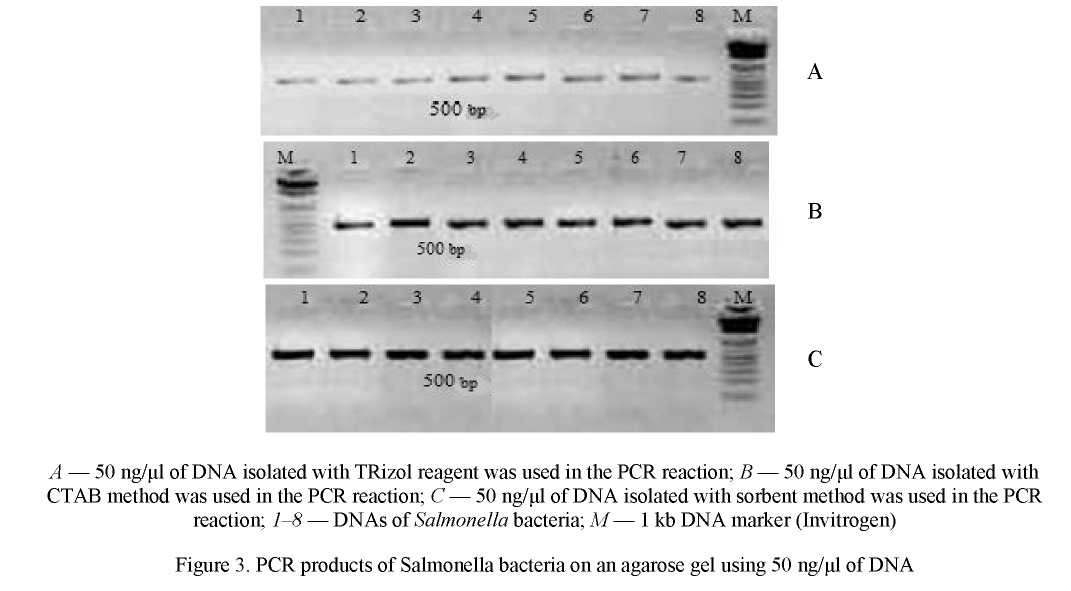
As can be seen in Figure 3, optimal results were obtained when PCR was performed using DNA isolated by the sorbent method.
The major criterion in methods of the DNA extraction is that nucleic acid should be the utmostly purified from cellular DNA and protein impurities. The extracted genome DNA should be unfragmented since it is a matrix for the synthesis of the specific product [9]. Such a result is ensured by the sorbent method using silica particles. When this method has used the proteins and cellular components are removed and the utmost purified DNA is left. The advantages of the DNA extraction method with the help of silica particles are as follows: loss minimization in the course of the DNA extraction; lower risk of cross-contamination owing to the linkage of all nucleic acid to the sorbent; high purity of the final product [10]. In contrast to other methods, this one does not require the usage of toxic reagents.
Effectiveness of the process of the DNA extraction is a determining factor in the course of work with a small amount of the material or with the specimens containing a considerable quantity of inhibitors that may result in lower diagnostic sensitivity of the test and inhibition of the amplification process [11]. The properly selected method of the DNA extraction allows achieving the most accurate result of the polymerase chain reaction.
Conclusion
On the basis of the implemented study, one can conclude that the sorbent method using silica particles is the most suitable method for DNA extraction from Salmonella. In the future, this method may be used to kit up the test system for Salmonella identification and genotyping.
The work was carried out within the project of grant financing «Genotyping pathogenic microorganisms in the food raw material and foodstuff realized in the markets and supermarkets of the Republic of Kazakhstan, development of recommendations aimed at reducing the risk of morbidity among the children of preschool and school age», years 2018–2020, No. AP05131147.
References
- Tokarev, V.A., Gudkova, E.I., & Adarchenko, A.A. (1999). Varianty monitorinha hospitalnykh shtammov VBI salmonelleznoi etiolohii [Variants of monitoring hospital strains of nosocomial infections, Salmonella etiology]. Proceedings of Nosocomial infections — problems of epidemiology, clinic, diagnosis, treatment, and prevention. Rossiiskaia nauchno-prakticheskaia konferentsiia — Russian scientific-practical conference (pp. 236-237) [in Russian].
- Lim, J.K., Gunther, N.W., Zhao, H., Johnson, D.E., Keay, S.K., & Mobley, H.L.T. (1998). In Vivo Phase Variation of Escherichia coli Type 1 Fimbrial Genes in Women with Urinary Tract Infection. Infection and immunity, 66, 7, 3303-3310.
- Abzhalieva, A.B. (2017). Usovershenstvovanie metodov identifikatsii pishchevykh patohenov v produktakh zhivotnoho pro- iskhozhdeniia [Improving the identification of foodborne pathogens in animal products]. Extended abstract of doctor's thesis. Almaty [in Russian].
- Nurgalieva, M.T., Zhanserkenova, O.O., Usenbekov, E.S., Kasymbekova, Sh.N., & Smagulov, A.K. (2016). Metody vydeleniia nukleinovykh kislot dlia provedeniia polimeraznoi tsepnoi reaktsii v rezhime realnoho vremeni [Nucleic acid isolation methods for real-time polymerase chain reaction]. Izdenister, natizheler — Research results, 3, 71, 68-72 [in Russian].
- Mamaev, M.A. (2000). Razrabotka uskorennykh sposobov DNK-diahnostiki dlia opredeleniia bakterii v miase i miasoproduk- takh [Development of accelerated methods of DNA diagnostics for the detection of bacteria in meat and meat products]. Extended abstract of candidate's thesis. Moscow [in Russian].
- Herrera-Leo, S., McQuiston, J.R., & Usera, M.A. (2004). Multiplex PCR for Distinguishing the Most Common Phase-1 Flagellar Antigens of Salmonella spp. J. Clin. Microbiol, 42, 2581–2586.
- Scott O. Rogers, Arnold J. Bendich (1985). Extraction of DNA from milligram amounts of fresh, herbarium and mummified plant tissues. Plant Molecular Biology, 5, 69–76.
- Kapustin, D.V., Prostiakova, A.I., & Alekseev, Ya.I. (2014). Vysokoeffektivnyi metod odnostadiinoho vydeleniia DNK dlia PTsR-diahnostiki Mycobacterium tuberculosis [Highly effective single-stage DNA extraction method for PCR diagnostics of Mycobacterium tuberculosis]. Acta naturae, 2, 21, 52–57 [in Russian].
- Kirkimbaeva, Zh.S., Ermagambetova, S.E., & Biiashev, K.B. (2016). Sravnitelnoe izuchenie effektivnosti razlichnykh metodov vydeleniia DNK leptospir [A comparative study of the effectiveness of various methods of DNA isolation leptospiri]. Izdenister, nətizheler — Research results, 4, 72, 50–54 [in Russian].
- Aukenov, N.E., Masabaeva, M.R., & Khasanova, U.U. (2014). Vydelenie i ochistka nukleinovykh kislot. Sostoianie problemy na sovremennom etape [Isolation and purification of nucleic acids. The state of the problem at the present stage]. Nauka i zdra- vookhranenie — Science and Health, 1, 51–53 [in Russian].
- Kataloh saita «DNK-Tekhnolohiia» [Site Catalog «DNA-Technology»]. docplayer.ru. Retrieved from_ https://docplayer.ru/ 45703435-I-ekspress-metod-vydeleniya-dnk-komplekt-reagentov-dlya-vydeleniya-dnk-proba-rapid-4-komplekt-reagentov-dlya- vydeleniya-dnk-proba-rapid-genetika-5.htmljin Russian].