At present the traditional methods of identifying microorganisms are replaced by express methods, the mass spectrometric method using MALDI-TOF MS allows to reliably identify a variety of microorganisms in a short time, which is an indisputable advantage in work and allows to quickly identify many microorganisms in quickly. Classical methods for the identification of lactic acid bacteria based on their cultivation require a long time for their implementation. The advent of matrix laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) technology made significant changes in the workflows for the study of lactic acid bacteria, unmatched in speed characteristics. This article presents the study of the morphological, cultural properties, acid-forming ability, antibiotic sensitivity lactic acid bacteria of isolated from ... products (ayran, koumiss, ashykan kozhe, kurt, suzbe, cottage cheese) produced in different districts of the Karaganda region. The species identification of the isolated strains of lactic acid bacteria was carried out using a MALDI-TOF mass spectrometer. The studies carried out on cultural and morphological characters indicate that they belong to the genus Lactobacillus, Lacticaseibacillus, Lactiplantibacillus, Limosilactobacillus. As a result of identification on the mass spectrometer of the presented samples L. acidophilus (2 strains), L. delbrueckii subsp. bulgaricum (2 strains), L. rhamnosus (7 strains), L. plantarum (2 strains), L. paracasei (11 strains), L. fermentum (2 strains) were revealed. According to the Score values, the results indicate the accuracy of the identification.
Introduction
A significant disadvantage of the widely used traditional microbiological methods is the long time to obtain the result, which is due to the physiology of microorganisms that requires time for their growth. Modern microbiology requires new fast and accurate methods for the identification of microorganisms.
The advent of matrix laser desorption ionization time-of-flight mass spectrometry technology has made significant changes in the workflow of microbiology laboratories, virtually unmatched in the speed and accuracy of microbial identification. This method, which is gradually becoming the standard for modern laboratories, makes it possible to reduce the identification time of pure cultures to several minutes [1].
The method is based on the extraction of peptides and proteins from cells of microorganisms and works according to the principle (molecular «fingerprint»), which is compared with the reference spectra in the MALDI bio typer database [2].
The MALDI TOF mass spectrometry method allows not only to identify a microorganism but also, in some cases, obtain a unique set of ribosomal proteins (fingerprints) for each of the strains under study, which opens up great opportunities and prospects for studying strain characteristics [3].
Bacteria of the genus Lactobacillus has always attracted attention of scientists and researchers around the world due to its great practical value.
The genus Lactobacillus includes more than 261 species (as of March 2020), which are very diverse at the phenotypic and genotypic levels. The genus was reclassified based on phylogenetic studies of the main genome, taking into account the physiological and ecological criteria: the corrected genus Lactobacillus was retained and 23 new genera were added [4].
Classical methods for the identification of lactic acid bacilli based on their cultivation require a long time for their implementation. The advent of matrix laser desorption ionization time-of-flight mass spectrometry (MALDI-TOF MS) technology made significant changes in the workflows for the study of lactic acid bacilli, unmatched in speed characteristics.
Objective: identification and study of the morphological and cultural properties, acid formation activityand the resistance to antimicrobial preparation of lactic acid bacilli isolated from lactic acid products Karaganda region by time-of-flight MALDI-TOF mass spectrometry (in vitro study).
Methodology
- 3 samples of lactic acid products were used for the study (ayran, koumiss, ashykankozhe, kurt, suzbe, cottage cheese) produced in different areas of the Karaganda region.
Sampling: sampling was carried out in accordance with the rules of asepsis. Before analyzing samples for the presence of probiotic cultures liquid dairy products (ayran, koumiss, ashykan kozhe) were stirred in a circular motion, avoiding active shaking to prevent air saturation. Cottage cheese, suzbe — were mixed by shaking.
- liquid dairy products were taken with a sterile pipette (10 ml) and placed in sterile test tubes.
- a pasty dairy product (suzbe) was taken with a sterile spatula (10 g) and placed into sterile test tubes.
- 10 g of kurt and cottage cheese were weighed on a Petri dish, transferred to a sterile porcelain mortar with a pestle, and thoroughly ground [5–6].
Study of morpho-cultural properties, mobility, and catalase test: Before sowing, a tenfold dilution was prepared from each product in sterile saline, followed by sowing on Petri dishes with agar MRS medium. The Petri dishes were cultured at 37°С for 2 days in anaerobic conditions.
After Petri dishes incubation, 42 strains of lactic acid bacilli were isolated from the milk product (ayran, koumiss, ashykankozhe, kurt, suzbe, cottage cheese), of which 26 isolated colonies were typical of lactic acid bacilli, microscoped (Gram stain) and seeded on MRS broth.
After 2 days of incubation control, smears were made from all tubes with broth, after which the isolate individual colonies by ten-fold dilution method, followed by seeding on Petri dishes with agarised MRS medium. Crops were incubated in anaerobic conditions at 371оС temperature for 48 hours [7–8].
After incubation, isolated colonies were determined by relation to Gram stain, mobility, presence of catalase and identified on MALDI BioTyper.
To determine the ratio of the isolated strains to Gram stain, smears were prepared from the colonies, stained according to the Gram method and microscoped. We took photos using a ToupcamTM USB digital eyepiece camera Industrial digital camera, 14 megapixels.
Catalase activity test: the catalase activity of the cultures was determined on the ability of catalase to decompose hydrogen peroxide with the release of gas bubbles. The reaction was set up with a daily culture cooled to room temperature on a sterile glass slide. An isolated colony taken from the surface of the nutrient medium was ground on glass and a drop of 3 % hydrogen peroxide solution was applied with a pipette. If after 30–60 s gas bubbles appear on the glass, then the reaction results are considered positive. A control sample was placed in parallel. The mobility of the isolated cultures was determined by «the crushed drop» [9].
The grown crops were identified using the MALDI BioTyper. The samples were prepared as follows: a fresh single colony was directly transferred onto a polished steel MSP 96 target (Bruker Daltonik) and dried; coated with 1µl of a saturated solution of a-cyano-4-hydroxy-cinnamic acid (HCCA) matrix solution in 50 % acetonitrile-2.5 % trifluoroacetic acid (Bruker Daltonik) and dried at room temperature, after that the sample was subjected to analysis of the bacteria protein using MALDI-TOF MS system [10].
The identification reliability criteria were judged by the following score values: 2,300 — 3,000 — highly probable species identification; 2,000–2,299 — reliable genus identification, probable species identification; 1,700 — 1,999 — probable identification of the genus; 0–1.699 — identification failed.
Determination of acid formation activity. Two tubes of each culture were removed and put into a refrigerator for rapid cooling to prevent further acid production. Then, 10 ml of culture liquid was added to glass flasks, and phenolphthalein was added as an indicator 1 drop.
The total acidity was determined by titration of decinormal alkali NaOH, which was added dropwise from the burette to the retorts with the poured culture liquid until a stable pink stain appeared. The amount of decinormal alkali that was used for titration corresponds to the amount of decinormal acid produced in 10 ml of culture liquid [11].
Data of acid formation activity, expressed in degrees Turner /°Т/, was calculated by the formula:
° T = a × k × 10,
where a is the number of milliliters of 0,1М caustic soda solution to be titrated; k is a correction to the titer of 0,1М caustic soda solution; 10 is a correction factor for the mass of the analyzed sample.
The sensitivity of probiotic bacteria to antibiotics was determined by disk-diffusion test. From the test cultures, the suspensions conforming to the optical turbidity standard of 5 units (with a microbial body content of about 1,5х108 CFU/ml) were prepared, 1 сm3 of the culture suspension was applied to each agar medium dish, uniformly distributed over the surface by lawn method and slightly dried in laminar flow. Further, the antibiotic discs of 5 pieces were applied to the surface of the nutrient medium seeded with a suspension of lactic acid bacilli cells. Inoculated plates with discs were incubated at 37 ± 1 °C for 48 h. The antibiotic graph was formed by the diameter of the growth retardation zone of microorganisms. The study was performed in triplicate and the results were expressed as arithmetic mean.
Results and Discussion
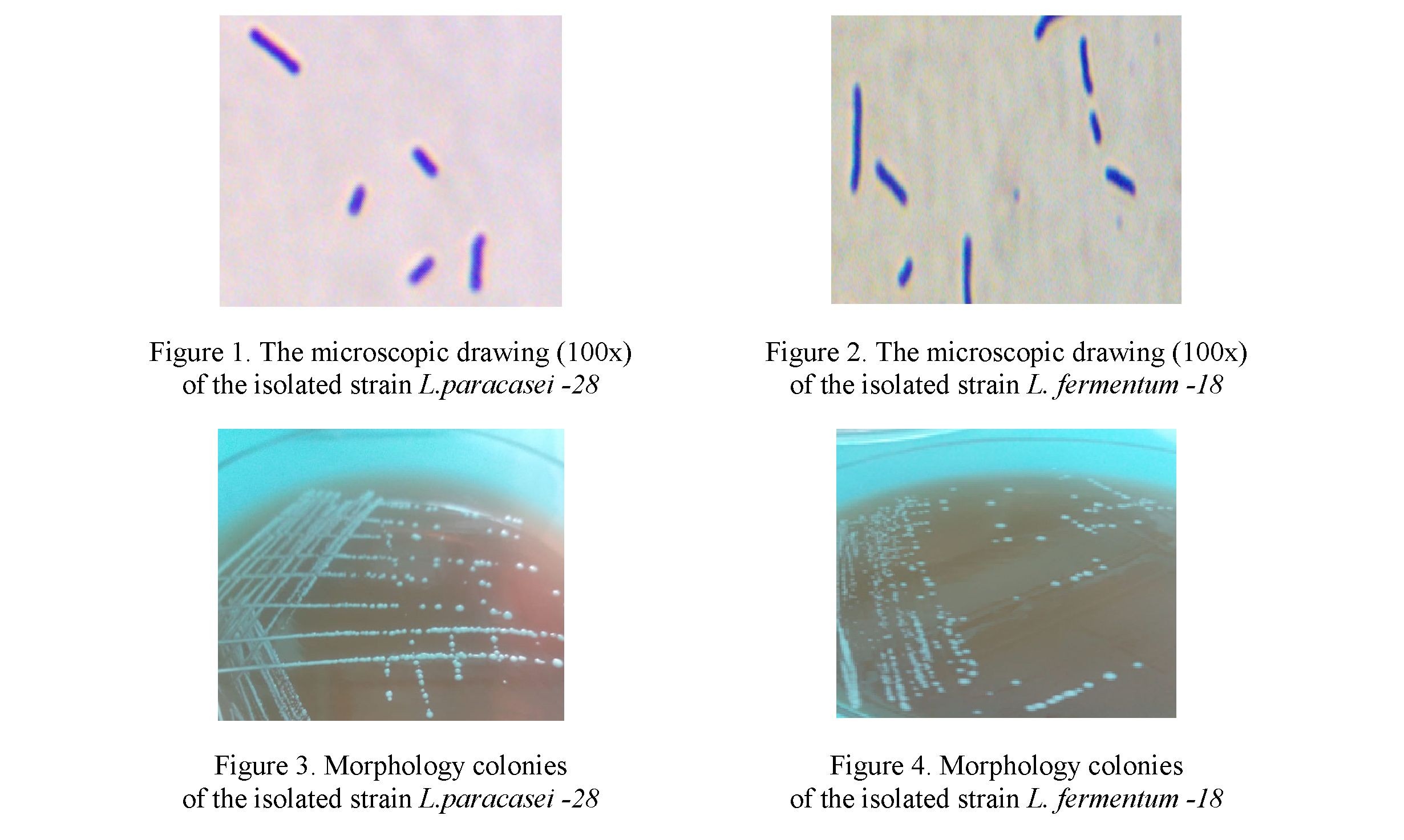
The studies carried out on cultural and morphological characters indicate their belonging to the genus Lactobacillus, Lacticaseibacillus, Lactiplantibacillus, Limosilactobacillus. All isolated strains are grampositive rods, cells are located singly, in pairs, in the form of a chain, motionless, spores do not form (Fig. 1, 2), catalase-negative (lactic acid bacilli do not have catalase, therefore, no gas formation was observed in the sample with hydrogen peroxide). Colony morphology (Figure 3, 4) of isolated strains of lactic acid bacilli are identified on solid nutrient medium MRS agar-1.
As a result of identification on the mass spectrometer of the presented samples it was L. acidophilus (2 strains), L.delbrueckiisubsp. bulgaricum (2 strains), L.rhamnosus (7 strains), L. plantarum (2 strains), L.paracasei (11 strains), L. fermentum (2 strains) — only 26 strains were revealed.
According to the Score values, the results indicate the accuracy of identification: Score values 2.300–3.000 are observed in 7 strains; Score values 2.000–2.299 are recorded in 15 strains; Score values 1.700–1.999 — in 4 strains.
Acid formation activity is a normalized indicator of biological activity of lactic acid bacilli and accordingly a criterion for selection of lactic acid bacilli strains with high-active probiotic properties. The results obtained in the study (Figure 5, 6) show good acid-forming ability in most isolated lactic acid bacilli

strains. Strains of lactic acid bacilli, titrated acidity of which varies within 20–80 °T are considered inactive, 90–110° T — average activity, and 120 ° T and higher are considered highly active.

Figure 5. Acid formation activity of lactic acid bacilli selected strains — average activity
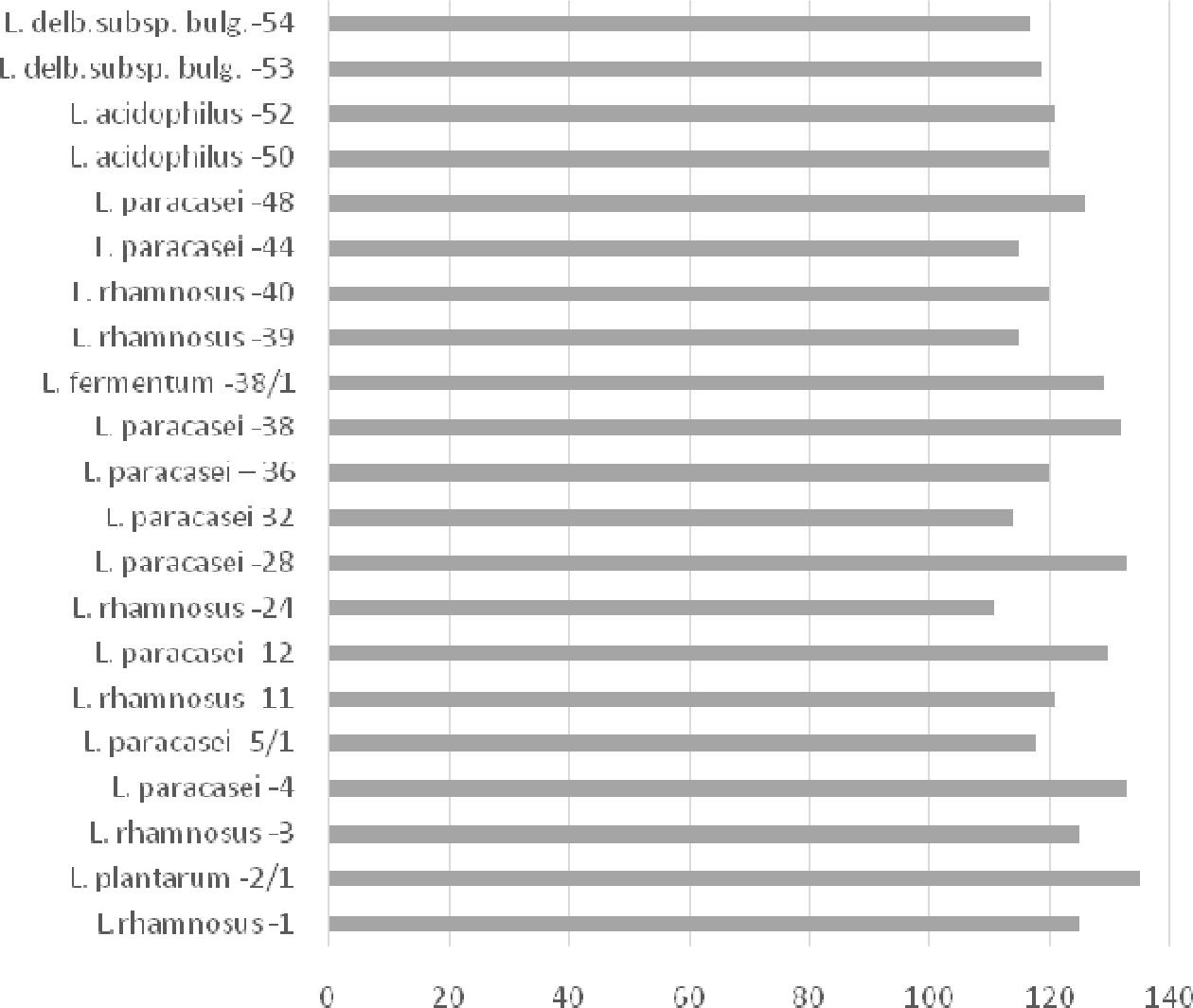
Figure 6. Acid formation activity of lactic acid bacilli selected strains — high activity
The resistance of bacteria to antimicrobial preparation is a characteristic feature of a particular strain of the microorganism and this should be taken into account when selecting cultures, products and preparations with probiotic properties used in biotechnology. In this regard, we have conducted studies to determine the spectrum of antibiotic resistance of isolated strains of lactic acid bacilli, to various most common antibiotics in medical practice.
The study found that 10 strains L.paracasei, 5 strains L. rhamnosus, 2 strains L. fermentum, 2 strains L. acidophilus, 1 strains L. delbr. subsp. bulg. are metronidazole (5 μg/disc) resistant the following strains are sensitive to antibiotic metronidazole L. paracasei — 36 (16 mm), L. plantarum -2/1- (8 mm), L. plantarum -5-(10 mm), L. rhamnosus -24 — (11 mm), L. plantarum -2/1 — (8 mm), L. delbr. subsp. bulg. -53 — (18 mm); 9 strains L.paracasei, 5 strains L. rhamnosus, 2 strains L. fermentum, 1 strains L. acidophilus, 1 strains L. delbr. subsp. bulg. showed resistance to сolistin (25 μg/disc), the following strains are sensitive to antibiotic сolistin L. paracasei — 4 (8 mm), L. paracasei — 36 (10 mm), L. plantarum -2/1- (9 mm),L. plantarum -5- (7 mm), L. rhamnosus -24 — (23 mm), L. plantarum -2/1- (9 mm), L. acidophilus -50(24 mm), L. delbr. subsp. bulg. — 54 — (12 mm).
Resistance was observed in 4 strains L.paracasei, 1 strains L. rhamnosus, 1 strains L. acidophilus to сefuroxime (30 μg/disc), sensitive to antibiotic сefuroxime the following strains L. paracasei — 4 (23 mm), L. paracasei -5/1- (23 mm), L. paracasei -12- (31 mm), L. paracasei — 36 (33mm), L. paracasei -38 - (19 mm), L. paracasei -44-(33 mm), L. paracasei -48 -(22 mm), L. rhamnosus -1-(33 mm), L. rhamnosus -2(23 mm), L. plantarum -2/1-(17 mm), L. rhamnosus -3-(17 mm), L. plantarum -5-(20 mm), L. rhamnosus - 11-(30 mm), L. fermentum -18-(18 mm), L. rhamnosus -24-(31 mm), L. rhamnosus -40-(45 mm), L. plantarum -2/1-(17 mm), L. fermentum -38/1-(27 mm), L. acidophilus -50-(34 mm), L. acidophilus -52(17 mm), L. delbr. subsp. bulg. -53-(23 mm), L. delbr. subsp. bulg. — 54-(33 mm).
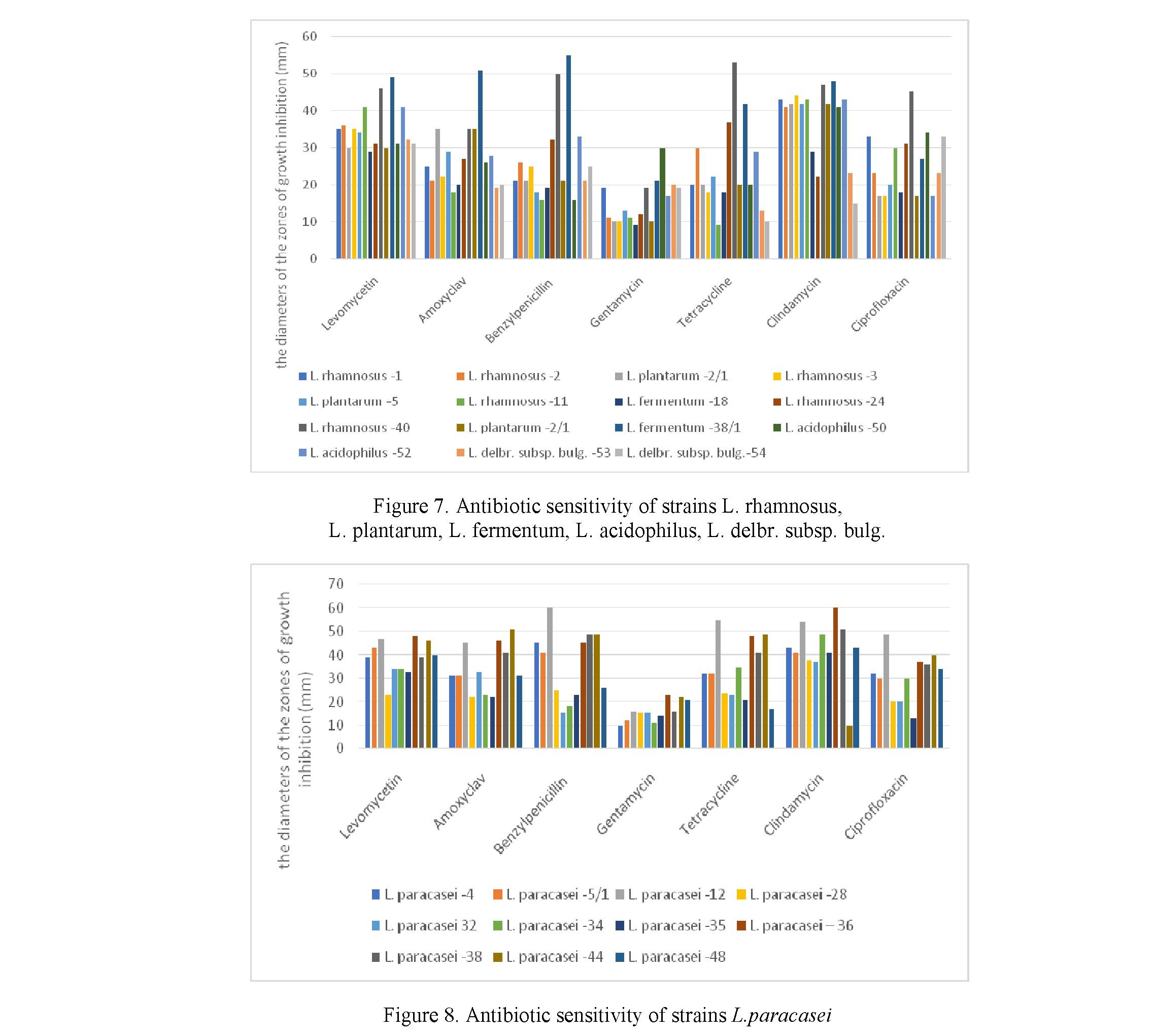 Most strains of lactic acid bacilli were sensitive to the following antibiotics (Figure 7, 8): levomycetin (10 μg/disc), amoxyclav (10 μg/disc), benzylpenicillin (10 μg/disc), gentamycin (10μg/disc), tetracycline (10 μg/disc), clindamycin (10 μg/disc), ciprofloxacin (30 μg/disc).
Most strains of lactic acid bacilli were sensitive to the following antibiotics (Figure 7, 8): levomycetin (10 μg/disc), amoxyclav (10 μg/disc), benzylpenicillin (10 μg/disc), gentamycin (10μg/disc), tetracycline (10 μg/disc), clindamycin (10 μg/disc), ciprofloxacin (30 μg/disc).
Conclusions
As a result of the studies carried out it can be concluded that it is impossible to judge with confidence the species homogeneity or heterogeneity of the culture by morpho-cultural characters, since the shape and size of microbial cells, the shape of the lactic acid bacilli colony varies markedly. Factors such as culture conditions, bacterial growth phases, and others inhibit the work of the researcher. Our research has shown that using MALDI-TOF-MS analyzes has become a part of the daily practical work of a microbiologist in identifying lactic acid bacilli, and even surpasses the (classical) methods of microbiological identification previously used by us in terms of speed and reliability.
According to literary data [12], in vitro lactic acid bacilli sensitivity tests are still poorly standardized. The evaluation of antibiotic sensitivity of these bacteria is extremely difficult, because the size of the zones recommended for other bacteria are not applicable to them. To these reasons are added specific conditions of cultivation: enriched medium, complex composition of the atmosphere, prolonged incubation.
Domestic microbiologists K.Kh. Almagambetov, I.S. Savitskaya et.al. do not give the clear criteria for the classification of strains of lactic acid bacilli to sensitive or resistant [13]. Therefore, empirical concentrations in the antibiotic disc (μg/disc) that delayed the growth of at least one of the strains studied were selected as boundary minimum inhibitory concentrations (MIC).
References
- Popov, D.A., Ovseenko, S.T., & Vostrikova, T.Yu. (2016). Primenenie metoda MALDI-TOF MS v sovremennoi mikrobiologicheskoi laboratorii [Application of the MALDI-TOF MS method in a modern microbiological laboratory]. Spetsvypusk LABORATORIIA LPU — Special LABORATORY to the clinic, 8, 53–56 [in Russian].
- Lavrinenko, A.V., Azizov, I.S. & Kolesnichenko, S.I. (2016). Ispolzovanie metodov MALDI-TOF mass-spektrometrii i pitatelnoi sredy Chamada Candida dlia identifikatsii gribov roda Candida [Using the methods of MALDI-TOF mass spectrometry and nourishing environment Chamada Candida for identification of fungi of the genus Candida]. Uspekhi meditsinskoi mikologii — Advances in medical mycology, 74, 75 [in Russian].
- Tochilina, A.G., Belova, I.V., Soloveva, I.V., Novikova, N.A., Ivanova, T.P. & Zhirnov, V.A. (2015). Izuchenie biologicheskikh svoistv shtammov roda Lactobacillus [Study of biological properties of Lactobacillus strains]. Sovremennye problemy nauki i obrazovaniia — Modern problems of science and education, 5 [in Russian].
- Zheng, J., Wittouck, S., Salvetti, E., Franz, C., Harris, H., Mattarelli, P., O’Toole, P. W., Vandamme, B.P., Walter, J., Watanabe, K., Wuyts, S., Felis, G.E., Gänzle, M.G. & Lebeer S. (2020). A taxonomic note on the genus Lactobacillus: Description of 23 novel genera, emended description of the genus Lactobacillus Beijerinck 1901, and union of Lactobacillaceae and Leuconostocaceae. International Journal of Systematic and Evolutionary Microbiology, 70, 2782–2858.
- Mezhgosudarstvennyi standart. Moloko i molochnaia produktsiia. Metody mikrobiologicheskogo analiza [Interstate standard milk and dairy products. Methods of microbiological analysis]. (2014) [in Russian].
- HOST R–56139–2014. Produkty pishchevye spetsializirovannye i funktsionalnye. Metody opredeleniia i podscheta probioticheskikh mikroorganizmov [Specialized and functional food products. Methods for determining and counting probiotic microorganisms]. (2016) [in Russian].
- Labinskaia, A.S., Blinkova, L.P. & Eshchina, A.S. (2004). Obshchaia i sanitarnaia mikrobiologiia s tekhnikoi mikrobiologicheskikh issledovanii [General and sanitary microbiology with microbiological research techniques]. Moscow: Meditsina [in Russian].
- Yarullina, D.R. & Fakhrullin, R.F. (2014). Bakterii roda Lactobacillus: obshchaia kharakteristika i metody raboty s nimi [Bacteria of the genus Lactobacillus: general characteristics and methods of working with them]. Kazan: Kazan University [in Russian].
- Sboichakov, V.B. & Karapats, M.M. (2014). Mikrobiologiia, virusologiia i immunologiia: rukovodstvo k laboratornym zaniatiiam [Microbiology, Virology and Immunology: a guide to laboratory classes]. Mocow: GEOTAR-Media [in Russian].
- Schulthess, B., Bloemberg, G.V., Zbinden, R., Böttger, E.C. & Hombach, M. (2014). Evaluation of the Bruker MALDI Biotyper for identification of Gram-positive rods: development of a diagnostic algorithm for the clinical laboratory. Journal of clinical microbiology, 52, 4, 1089–1097.
- Obshchaia farmakopeinaia statiia «Opredelenie spetsificheskoi aktivnosti probiotikov» [General Pharmacopoeia article «Determination of specific activity of probiotics»]. (2015). [in Russian].
- Bagdasarjan, A.S., Tokaev, Je.S., Nekrosav, E.A. & Olejnik, E.A. (2011). Antibiotikoustoichivost probioticheskih kultur, vkhodiashchikh v sostav sinbiotikov [Antibiotic resistance of probiotic cultures included in the synbiotic]. Izvestiia vuzov. Pishchevaia tekhnologiia — News of institutes of higher education. Food technology, 2–3, 102–104 [in Russian].
- Savickaja, I.S., Zhubanova, A.A., Kistaubaeva, A.S. & Bolekbaeva, A.B. (2012). Antibiotikorezistentnost laktobatsill- probiotikov [Antibiotic resistance of lactobacilli-probiotics]. Vestnik Kazakhskogo natsianalnogo universiteta. Seriia biologicheskaia — Bulletin of KazNU. Biology series, 56, 4, 222–227 [in Russian].